 |
 |
 |
 |
 |
 |
Horizontal Tabs
|
PDB ID |
Structure Details |
|
Structures SETDB1 Tudor domain in complex with fragments |
|
|
Structures SETDB1 Tudor domain in complex with fragments |
|
|
Structures SETDB1 Tudor domain in complex with fragments |
|
|
Structures SETDB1 Tudor domain in complex with fragments |
|
|
Structures SETDB1 Tudor domain in complex with fragments |
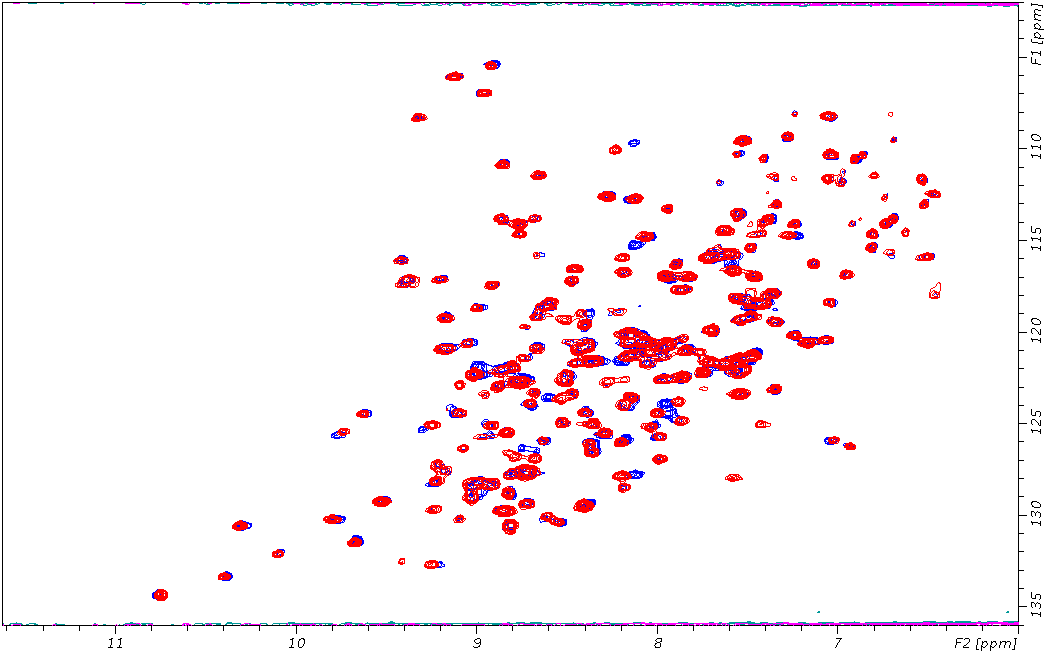
HSQC NMR of 10mM of MRT10241258a (blue) with SETDB1 and SETDB1 with DMSO (red)
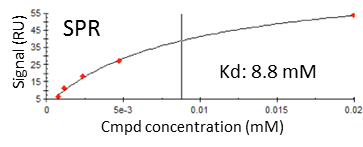
SPR data of one of the fragment (Chemical stucture not presented)
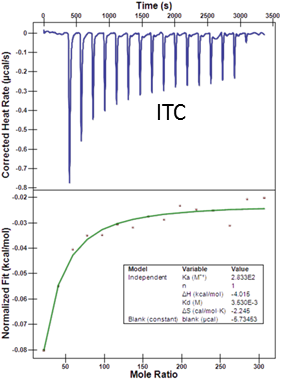
ITC data of one of the fragment (Chemical stucture not presented)
Experimental procedures
SPR Assay
Biotinylated SETDB1 captured onto a SA chip.
Buffer: 0.01 M HEPES pH 7.4, 0.15 M NaCl, 3 mM EDTA, 0.005% v/v Surfactant P20 5% DMSO Final Compound stock concentrations varied from 400 uM to 50 mM maintaining 5% DMSO final.
Flow rate: 25 µL/min, compound contact time: 30 seconds at each concentration. Compounds were tested using Single Cycle Kinetics
ITC Assay
Buffer: 20 mM HEPES pH 7.5, 250 mM NaCl
Temperature 25C
Cell: SETDB1 0.08 mM
Syringe: Fragments 50-100mM
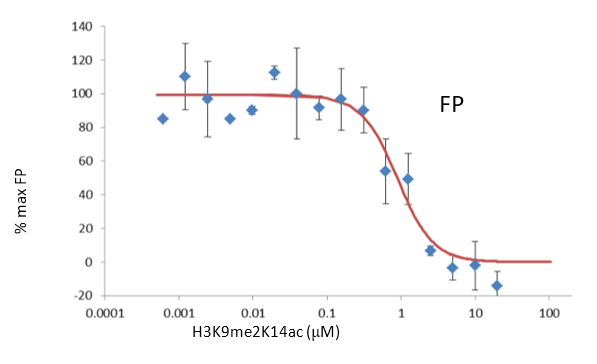
FP Assay
20 mM Hepes pH 7.5, 50 mM NaCl, 5 mM MgCl2, 0.01% TX100, 2 mM
DTT, containing 0.04 μM of 5' FITC- H3K9me2K14ac (1-25) and 5 μM SETDB1 at room temperature.
SETDB1 Tudor Protein Expression (used for crystallography, ITC, FP)
The expression construct for N-terminal His6-tagged SETDB1 Tudor domain (residues 197 – 403) subcloned into pET28a-MHL vector was transformed into BL21(DE3)-pRARE2 cells. The cultures were grown in Terrific Broth medium (TB) in the presence of 50 µg/mL kanamycin and 34 µg/mL chloramphenicol at 37 °C. When the OD600 reached 1.5, the overexpression of SETDB1 Tudor was induced by addition of isopropyl-1-thio-D-galactopyranoside (IPTG), final concentration 0.5 mM, and incubated overnight at 16 °C. Next day, the cells were harvested by centrifugation at 12,227 × g (10 min, 4C) and the cell pellets were flash frozen in liquid N2 and stored at -80 °C.
15N SETDB1 Tudor Protein Expression (used for 1H-15N TROSY NMR)
The expression construct for N-terminal His6-tagged SETDB1 Tudor domain (residues 197 – 403) subcloned into pET28a-MHL vector was transformed into BL21(DE3)-pRARE2 cells. The cultures were grown in auto-induction M9 minimal medium containing 1.2 g/l 15N-ammonium chloride (Sigma 299251) as the sole nitrogen source using an auto-induction method described in (Adelinda A. Yee, Anthony Semesi, Maite Garcia, and Cheryl H. Arrowsmith (2014) Screening Proteins for NMR Suitability. Wayne F. Anderson (ed.), Structural Genomics and Drug Discovery: Methods and Protocols, Methods in Molecular Biology, vol. 1140, DOI 10.1007/978-1-4939-0354-2_13.). The medium contained 50 µg/mL kanamycin and 34 µg/mL chloramphenicol at the whole cultivation was done at 37 °C. The cells were harvested by centrifugation at 12,227 × g (10 min, 4C) and the cell pellets were flash frozen in liquid N2 and stored at -80 °C.
Biotinylated SETDB1 Tudor Protein Expression (used for SPR)
The expression construct was N-terminally biotinylated SETDB1 Tudor domain (residues 197 – 403) subcloned into pET28BIOH_LIC vector. Details of the expression and purification can be obtained from Ashley or Mani.
Purification of SETDB1 Tudor (same procedure for unlabeled and 15N-labeled, different MW)
Purification procedure was based on the Materials and Methods section of
The cell pellets were thawed and resuspended (1 g of cell pellet per 10 mL) in binding buffer (20 mM HEPES pH 7.5, 500 mM NaCl, 5 mM imidazole, 0.5 mM TCEP, 5% glycerol) with protease inhibitor (0.1 mM phenylmethyl sulfonyl fluoride, PMSF). The cell suspension was supplemented with 0.5% (w/v) CHAPS, 5 µl of benzonase (EMD Millipore, cat. no. 70746), protease inhibitor cocktail (Roche) and the cells were sonicated on ice for 5 min total (10 s pulses with 5s interruptions). The lysate was clarified by centrifugation at 20,000 × g, 4 °C, 60 min and the resulting supernatant was filtered through 0.45 µm filter and applied onto 5 mL HisTrap HP column (GE). The column was washed with 10 CV of wash buffer (20 mM HEPES, pH 7.5, 500 mM NaCl, 40 mM imidazole, 0.5 mM TCEP, and 5% glycerol) and the protein was eluted using elution buffer (20 mM HEPES, pH 7.5, 500 mM NaCl, 250 mM imidazole, 0.5 mM TCEP, and 5% glycerol). Next the protein was applied on 26/60 Superdex 200 (GE) column equilibrated with 20 mM HEPES, pH 7.5, 150 mM NaCl, 0.5 mM TCEP at flow rate 2 ml/min. As the final purification step, the protein was purified to homogeneity by ion-exchange chromatography on Source 30S column (10x10) (GE), equilibrated with 20 mM HEPES, pH 7.4, and eluted with linear gradient of NaCl up to 500 mM concentration (20CV). Purification yield was 1 mg of the protein per 1L of culture. The MW (26293.2 Da for unlabeled and 26 616.2 Da for the 15N labeled) of the purified construct was confirmed by LC/MSD TOF (Agilent).
BIOH-SETDB1 Expression (used for SPR experiments)
The expression construct was N-terminally biotinylated SETDB1 Tudor domain (residues 197 – 403) subcloned into pET28BIOH_LIC vector. Growth was done as usual but supplemented with 1 mg/ml of biotin.
Purification of BIOH-SETDB1 Expression (used for SPR experiments)
1. Preparation of Solutions
IMAC lysis buffer: 50mM Tris-HCl, 300 mM NaCl, 5% glycerol, 10 mM imidazole, 0.5 mM TCEP, 0.05% Triton X100 pH 8.0
IMAC wash 1 buffer: 50mM Tris-HCl, 300 mM NaCl, 5% glycerol, 10 mM imidazole, 0.5 mM TCEP, pH 8.0
IMAC wash 2 buffer: 50mM Tris-HCl, 300 mM NaCl, 5% glycerol, 20 mM imidazole, 0.5 mM TCEP, pH 8.0
IMAC elution buffer: 50mM Tris-HCl, 300 mM NaCl, 5% glycerol, 300 mM imidazole, 0.5 mM TCEP, pH 7.5
Gel filtration buffer: 20 mM HEPES, 300 mM NaCl, 5% glycerol, 0.5 mM TCEP, pH 7.5
2. Samples preparation
1). The frozen cell pellets are briefly thawed in warm water. Resuspend the cell pellets by adding Lysis buffer to final volume 100 mL/(per liter culture), Freshly add PMSF and Benzamidin to a working concentration of 1mM and add Benzonase to 5U/ml.
2). Sonication with program nr 1 (5s/7s, 10 min, ~100W) for lysate from 2 L cultivation.
3). The sonicated lysates are centrifuged for 60 min at 16000 rpm in the JLA 16.250 rotor. The soluble fraction is decanted and filtered through 0.45mm filters.
3. Purification
The 5 mL chelating columns and gel filtration columns were equilibrated with IMAC wash 1 buffer and Gel filtration buffer, respectively.
Purification programs:
Chelating 5ml GFS75 2nd wash 20CV level and slope
Chelating 5ml GFS200 2nd wash 20CV level and slope
- Filtered samples are loaded to the IMAC columns on position 1-4 from sample inlets S1-S4
- IMAC columns are washed by 10 CV of IMAC wash 1 buffer and 10 CV of IMAC wash 2 buffer.
- IMAC columns are eluted by 5 CV IMAC elution buffer, and major peak is loaded to gel filtration columns on position 5.
- The fractions from gel filtration column are checked by SDS-PAGE and pool the pure fractions together. The antigens are further validated by Mass spectrum.
- The pure antigen samples are diluted or concentrated to 1 mg/ml and make 100 uL aliquots. The aliquots are flash freezed by liquid nitrogen and ship to binders lab with dry ice.
- Rodriguez-Paredes, M., Martinez de Paz, A., Simo-Riudalbas, L., Sayols, S., Moutinho, C., Moran, S., Villanueva, A., Vazquez-Cedeira, M., Lazo, P. A., Carneiro, F., Moura, C. S., Vieira, J., Teixeira, M. R., and Esteller, M. (2014) Gene amplification of the histone methyltransferase SETDB1 contributes to human lung tumorigenesis. Oncogene 33, 2807-2813
- Shah, M. A., Denton, E. L., Arrowsmith, C. H., Lupien, M., and Schapira, M. (2014) A global assessment of cancer genomic alterations in epigenetic mechanisms. Epigenetics Chromatin 7, 29
- Liu, N., Zhang, Z., Wu, H., Jiang, Y., Meng, L., Xiong, J., Zhao, Z., Zhou, X., Li, J., Li, H., Zheng, Y., Chen, S., Cai, T., Gao, S., and Zhu, B. (2015) Recognition of H3K9 methylation by GLP is required for efficient establishment of H3K9 methylation, rapid target gene repression, and mouse viability. Genes & development 29, 379-393
- Jurkowska, R. Z., Qin, S., Kungulovski, G., Tempel, W., Liu, Y., Bashtrykov, P., Stiefelmaier, J., Jurkowski, T. P., Kudithipudi, S., Weirich, S., Tamas, R., Wu, H., Dombrovski, L., Loppnau, P., Reinhardt, R., Min, J., and Jeltsch, A. (2017) H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1. Nature communications 8, 2057
We respectfully request that this document is cited using the DOI value as given above if the content is used in your work.
Please wait whilst the interactive viewer is loaded!
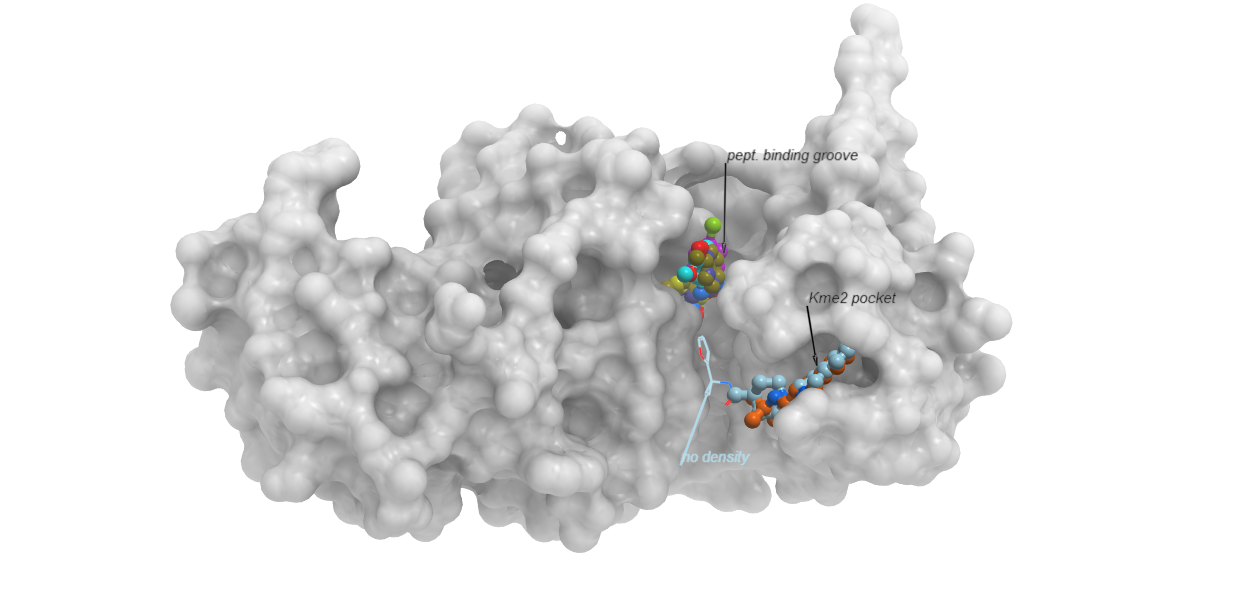
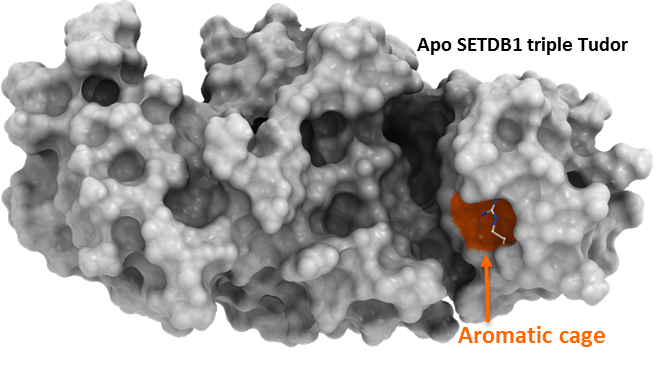
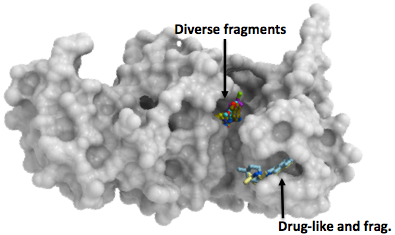
SETDB1 triple Tudor domain in complex with compounds occupying the methyl-lysine binding-pocket and the peptide-binding groove. (PDBs 5KH6, 5KCO, 5KCH, 5KE2, 5KE3).
