 |
 |
 |
 |
 |
 |
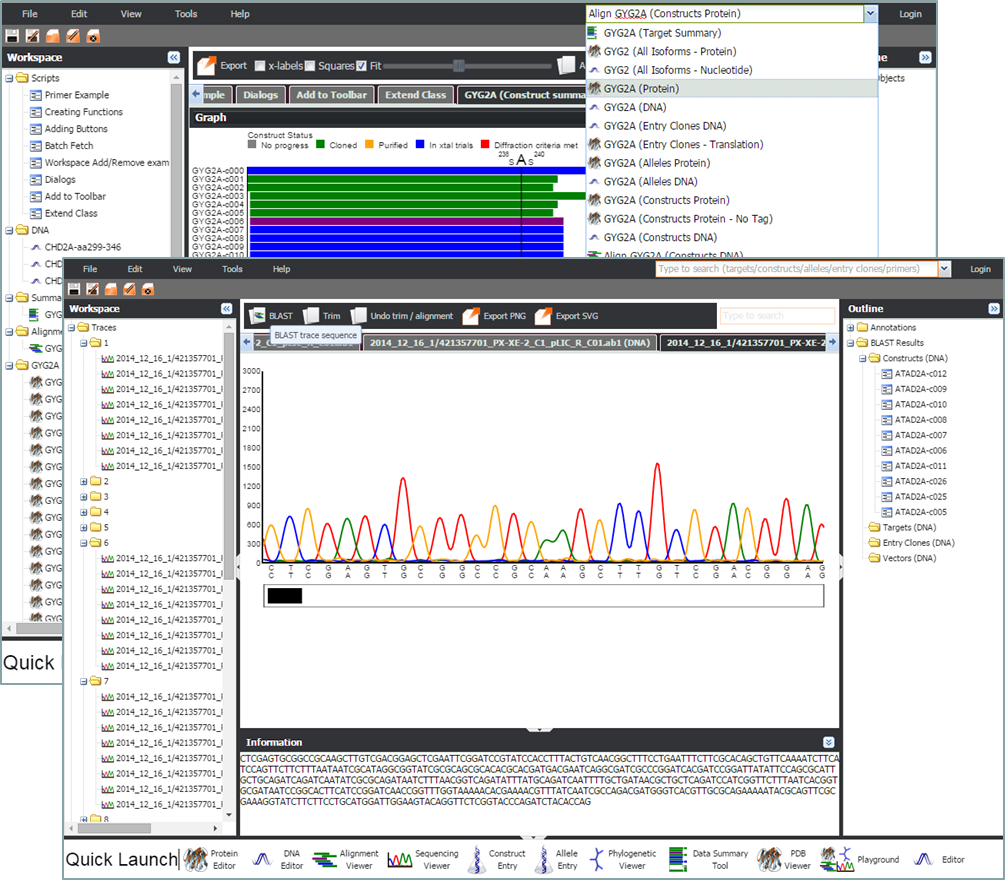 SATurn is an integrated solution for the storage & analysis of in-house: sequence collections, protein structures, compounds and glycans. SATurn is designed from the groundup to be easily extended for new data-types and tools.
SATurn is an integrated solution for the storage & analysis of in-house: sequence collections, protein structures, compounds and glycans. SATurn is designed from the groundup to be easily extended for new data-types and tools. The SATurn platform is not quite ready for general release. However if you are interested please email david.damerell@sgc.ox.ac.uk
Key Features
- Import / Export DNA & Protein sequences
- View common protein sequence annotations (i.e. secondary structure)
- Translate & Edit DNA sequences
- BLAST sequences against in-house databases
- Generate multiple sequence alignments and phylogenetic trees
- View protein structures
- View compound structures
- View ABI trace files (automatic alignment and mutation detection)
- View glycan structures
- Script editor built into interface
Platform
- HTML5 compliant (no plugins required)
- Available either as a standalone application or a hosted install
- Written in the Haxe programming language
- Extendable using either JavaScript or Haxe
- Will be licensed under the GPL
