|
 |
 |
 |
 |
 |
Modifications to DNA and histones are known to affect regulation gene expression in eukaryotes. Diseases such as cancer, inflammation, neurological and cardiovascular diseases can be related to aberrant histone modification patterns. Thus, there is great therapeutic interest in proteins that read, write, and erase chromatin marks since they may influence disease onset and progression. However, the identification of potent and selective inhibitors is challenging due to structural similarities between individual domains of the ‘epigenetic’ proteins.
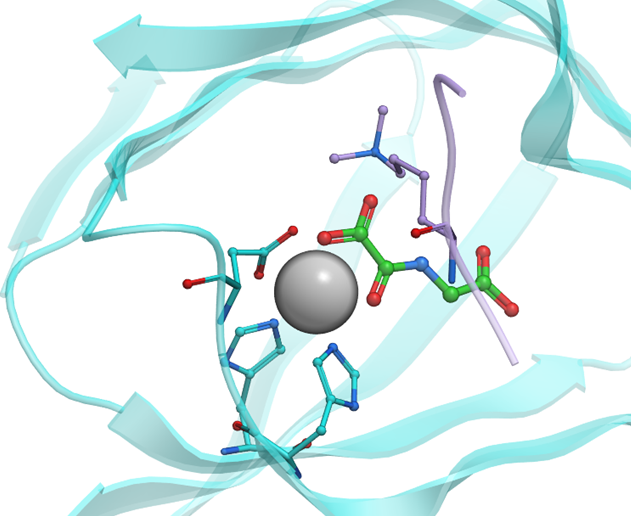
Histone methyltransferases and demethylases (KDMs) dynamically regulate histone lysine methylation levels. Removal of methyl groups from methylated lysines on histone tails is catalyzed by KDMs in a sequence- and methylation-state dependent manner. The two known families of KDMs are the flavin-dependent lysine specific demethylases (KDM1/LSD) and the JmjC-domain containing KDMs; this larger family of KDMs are Fe(II)- and 2-oxoglutarate (2OG)-dependent oxygenases (KDM2-7). Some KDM inhibitors have been reported (such as GSK-J1), however achieving selectivity remains a major challenge.