 |
 |
 |
 |
 |
 |
Chemical Probes
Chemical Probes
Potent, selective and cell-permeable inhibitors of protein function ("chemical probes") are valued reagents in both fundamental and applied biological research, and they are essential for the early stages of drug discovery by allowing preclinical target validation in both academic and industrial laboratories.
SGC chemical probes are open-access reagents that are meant to be used by the biomedical research community with no restrictions on use. To acquire a sample, follow the links on the webpage of your desired chemical probe. You can sort the table by clicking the column headers.
The SGC probes set is complemented by the donated probe set
| Name | Structures | Protein family | Specific Targets |
Control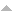 |
In Vivo Activity* | Added |
|---|---|---|---|---|---|---|
| BAY-069 |
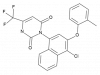 |
Amino acid transaminase | BCAT1, BCAT2 | BAY-771 | Yes | 09/20 |
| SGK3-PROTAC1 |
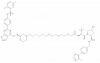 |
Kinase | SGK3 | cisSGK3-PROTAC1 | No | 01/24 |
| BI 653048 |
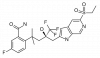 |
Nuclear hormone receptor | NR3C1 | BI-3047 | Yes | 11/18 |
| JNJ-42396302 |
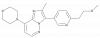 |
Phosphodiesterase | PDE10A | JNJ-40573663 | Yes | 09/21 |
| BAY-7598 |
 |
Protease | MMP12 | BAY-694 | Yes | 05/18 |
| I-BRD9 |
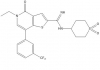 |
Bromodomains | BRD9 | No | 05/15 | |
| THPP-1 |
 |
Phosphodiesterase | PDE10A | THPP-1-NC | Yes | 06/19 |
| JNJ-9350 |
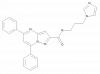 |
Oxidase | SMOX | JNJ-4545 | Yes | 09/22 |
| BAY-707 |
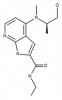 |
Hydrolase | NUDT1 | BAY-604 | Yes | 05/18 |
|
Bromosporine (BSP) (tool compound) |
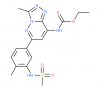 |
Bromodomains | pan-Bromodomain | No | 01/13 | |
| FHT-2344 |
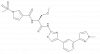 |
SWI/SNF | SMARCA2, SMARCA4 | FHT-5908 | Yes | 09/23 |
| ABT-546 |
 |
GPCR | EDNRA | A-545 | Yes | 05/18 |
| GSK789 |
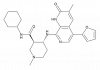 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | GSK791 | No | 02/21 |
| BI-1230 |
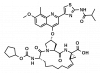 |
Protease | HCV NS3 | BI-1675 | Yes | 03/19 |
| BAY1125976 |
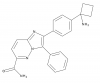 |
Kinase | AKT1/2 | BAY-940 | Yes | 11/21 |
| MRL-SYKi |
 |
Kinase | SYK | MRL-SYKi-NC | Yes | 05/18 |
| FS-694 |
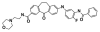 |
Kinase | MAPK14 | FM-743 | No | 01/20 |
| TP-030-1 |
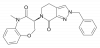 |
Kinase | RIPK1 | TP-030n | Yes | 09/20 |
| PFI-4 |
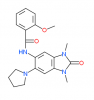 |
Bromodomains | BRPF1 | No | 12/14 | |
| BAY-277 |
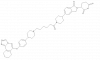 |
Aminopeptidase | METAP2 | BAY-8805 | Yes | 01/24 |
| BI-2545 |
 |
Phosphodiesterase | ENPP2 | BI-3017 | Yes | 11/18 |
| 8RK64 |
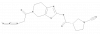 |
Regulator protein | UCHL1 | JYQ88 | Yes | 09/21 |
| BAY-474 |
 |
Kinase | MET | BAY-827 | Yes | 05/18 |
| BAY-549 |
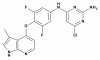 |
Kinase | ROCK1, ROCK2 | BAY-4900 | Yes | 06/19 |
| CCT369260 |
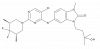 |
Transcription factor | BCL6 | CCT393732 | Yes | 09/22 |
| BI-9627 |
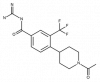 |
Solute carriers | SLC9A1 | BI-0054 | Yes | 05/18 |
| TP-060 |
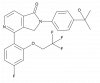 |
Glucosyltransferase | UGCG | TP-060n | Yes | 09/23 |
| KISS1-305 |
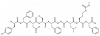 |
GPCR | KISS1R | KISS1-543 | Yes | 05/18 |
| BAY-3153 |
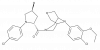 |
GPCR | CCR1 | BAY-173 | Yes | 02/21 |
| TP-021 |
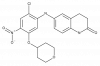 |
Transcription factor | BCL6 | TP-021n | Yes | 11/18 |
| Borussertib |
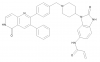 |
Kinase | AKT1, AKT2 | RL2578 | Yes | 10/21 |
| CRTH2 antagonist |
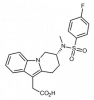 |
GPCR | PTGDR2 | CRTH2 negative control | Yes | 05/18 |
| SR-318 |
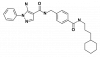 |
Kinase | MAPK14 | SR-321 | No | 01/20 |
| A-1211212 |
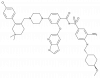 |
Regulator protein | BCL2 | A-1210227 | Yes | 11/19 |
| GSK-LSD1 |
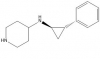 |
Lysine Demethylase | KDM1A/LSD1 | No | 03/14 | |
| Ogerin |
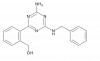 |
GPCR | GPR68 | ZINC32547799 | Yes | 06/20 |
| A-366 |
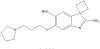 |
Methyltransferase | EHMT1/GLP, EHMT2/G9a | No | 06/13 | |
| BAY 1753011 |
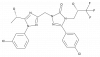 |
GPCR | AVPR1A, AVPR2 | BAY-2297 | Yes | 12/23 |
| BI-4394 |
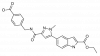 |
Protease | MMP13 | BI-4395 | Yes | 11/18 |
| BI-1942 |
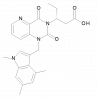 |
Protease | CMA1 | No | 06/21 | |
| MLi-2 |
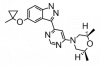 |
Kinase | LRRK2 | MLi-2-NC | Yes | 05/19 |
| JNJ-6204 |
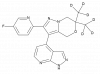 |
Kinase | CSNK1D/CSNK1E | JNJ-0293 | Yes | 09/22 |
| PF-05105679 |
 |
Ion channel | TRPM8 | PF-05257137 | Yes | 05/18 |
| JNJ-4355 |
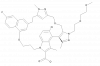 |
BCL2 protein family | MCL1 | JNJ-78732576 | Yes | 09/23 |
| ABT-100 |
 |
Transferase | FNTB | A-108 | Yes | 05/18 |
| BI 207127 |
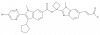 |
RNA polymerase | HCV NS5B | BI-7656 | Yes | 11/20 |
| BI-5524 |
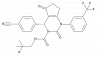 |
Chymotrypsin | ELANE | BI-5525 | Yes | 01/24 |
| BAY-784 |
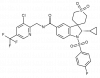 |
GPCR | GNRHR | BAY-786 | Yes | 11/18 |
| MSC-4381 |
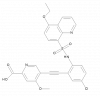 |
Monocarboxylate transporter | SLC16A3 | MSC-0516 | Yes | 09/21 |
| BI 665915 |
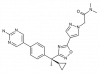 |
Lipoxygenase | ALOX5AP | BI-0153 | Yes | 05/18 |
| BAY-899 |
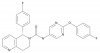 |
GPCR | LHCGR | BAY-897 | Yes | 01/20 |
| MSD-M1PAM |
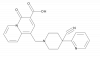 |
GPCR | CHRM1 | MSD-M1PAM-NC | Yes | 08/19 |
| UNC0642 |
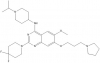 |
Methyltransferase | EHMT2/G9a, EHMT1/GLP | Yes | 08/12 | |
| BI-3812 |
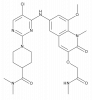 |
Transcription factor | BCL6 | BI-5273 | No | 11/23 |
| BAY-678 |
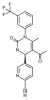 |
Protease | ELANE | BAY-677 | Yes | 05/18 |
| GSK973 |
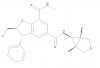 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | GSK943 | Yes | 02/21 |
| T3-CLK |
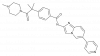 |
Kinase | CLK1, CLK2, CLK3, CLK4 | T3-CLK-N | No | 04/19 |
| TP-051 |
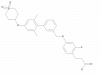 |
GPCR | FFAR1 | TP-051n | Yes | 09/22 |
| BI-1935 |
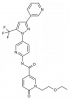 |
Hydrolase | EPHX2 | BI-2049 | Yes | 05/18 |
| SR-302 |
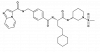 |
Kinase | DDR1, DDR2, MAPK11, MAPK14 | SR-301 | No | 06/20 |
| BAY-390 |
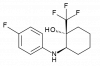 |
Ion channel | TRPA1 | BAY-9897 | Yes | 09/20 |
| BIBO3304 |
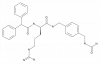 |
GPCR | NPY1R | BIBO3457 | Yes | 01/24 |
| BI-1950 |
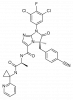 |
Integrin | ITGAL | BI-9446 | Yes | 11/18 |
| JNJ-54119936 |
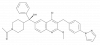 |
Nuclear hormone receptor | RORC | JNJ-53721590 | Yes | 09/21 |
| ERKi |
 |
Kinase | MAPK3, MAPK1 | ERKi-NC | Yes | 05/18 |
| NI-57 |
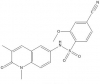 |
Bromodomains | BRPF1, BRD1, BRPF3 | No | 12/14 | |
| A-1155463 |
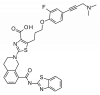 |
Regulator protein | BCL2L1 | A-1107969 | Yes | 11/19 |
| MSD-CYP11B2 |
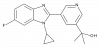 |
Cytochrome P450 | CYP11B2 | MSD-CYP11B2 Negative control | Yes | 06/19 |
| BAY 1816032 |
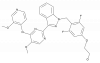 |
Kinase | BUB1 | BAY-283 | Yes | 10/22 |
| BAY-386 |
 |
GPCR | F2R | BAY-448 | Yes | 05/18 |
| GSK343 |
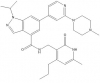 |
Methyltransferase | EZH2 | No | 08/12 | |
| JP3000 |
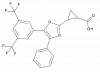 |
RXRA, RXRB, RXRG | JP3001 | 10/23 | ||
| T-26c |
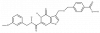 |
Protease | MMP13 | T-26f | Yes | 05/18 |
| GSK046 |
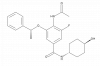 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | Yes | 02/21 | |
| A-192621 |
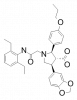 |
GPCR | EDNRB | A-1806262 | Yes | 03/19 |
| GNE-2256 |
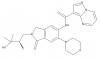 |
Kinase | IRAK4 | GNE-6689 | Yes | 11/21 |
| MRK-560 |
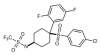 |
Protease | Gamma secretase complex | GSI-NC | Yes | 05/18 |
| BAY-3827 |
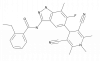 |
Kinase | PRKAA1, RPS6KA1 | BAY-974 | No | 01/20 |
| TP-030-2 |
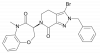 |
Kinase | RIPK1 | TP-030n | Yes | 09/20 |
|
IOX2 (requires >>> 1 microM for cell assays) |
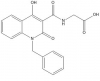 |
2OG | PHD2 | No | 02/12 | |
| VZ185 |
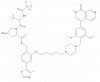 |
Bromodomain-containing protein family | BRD7, BRD9 | cisVZ185 | No | 01/24 |
| BI 639667 |
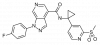 |
GPCR | CCR1 | BI-9307 | Yes | 11/18 |
| TP-040 |
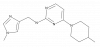 |
Hydrolase | OGA | TP-040n | Yes | 09/21 |
| BI 99179 |
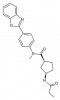 |
Synthase | FASN | BI 99990 | No | 05/18 |
| BI-9564 |
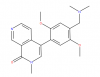 |
Bromodomains | BRD9, BRD7 | Yes | 04/15 | |
| PPTN |
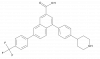 |
GPCR | P2RY14 | PPTN-NC | Yes | 06/19 |
| SAFit2 |
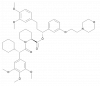 |
Isomerase | FKBP5 | ddSAFit2 | Yes | 09/22 |
| A-485 |
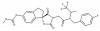 |
Transferase | EP300, CREBBP | A-486 | Yes | 05/18 |
| PFI-1 |
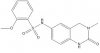 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | No | 11/11 | |
| BI-3231 |
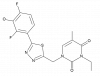 |
17β-hydroxysteroid dehydrogenases | HSD17B13 | BI-0955 | Yes | 09/23 |
| PF-04457845 |
 |
Hydrolase | FAAH | PF-04875474 | Yes | 05/18 |
| GSK778 |
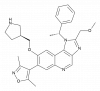 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | Yes | 02/21 | |
| TP-024 |
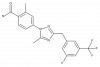 |
GPCR | GPR52 | TP-024n | Yes | 11/18 |
| JNJ-65355394 |
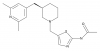 |
Hydrolase | OGA | JNJ-73924149 | Yes | 11/21 |
| MRL-650 |
 |
GPCR | CNR1 | MRL-CB1-NC | Yes | 05/18 |
| Skepinone-L |
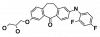 |
Kinase | MAPK14 | FM-743 | Yes | 01/20 |
| (R)-ZINC-3573 |
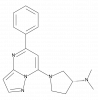 |
GPCR | MRGPRX2 | (S)-ZINC-3573 | No | 06/20 |
| BAZ2-ICR |
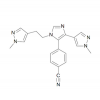 |
Bromodomains | BAZ2A, BAZ2B | Yes | 05/14 | |
| BAY-6096 |
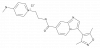 |
GPCR | ADRA2B | BAY-726 | Yes | 12/23 |
| BAY-885 |
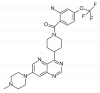 |
Kinase | MAPK7 | BAY-693 | No | 11/18 |
| A-446 |
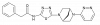 |
Hydrolase | GLS | A-426 | Yes | 07/21 |
| OF-1 |
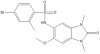 |
Bromodomains | BRPF1, BRD1, BRPF3 | No | 07/14 | |
| TP-008 |
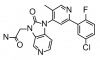 |
Kinase | ACVR1B, TGFBR1 | Al11 | No | 06/19 |
| JNJ-39758979 |
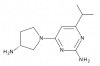 |
GPCR | HRH4 | JNJ-39668551 | Yes | 09/22 |
| TP-004 |
 |
Protease | METAP2 | TPn-004 | Yes | 05/18 |
| JNJ-42226314 |
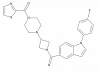 |
Serine hydrolase | MGLL | JNJ-8034 | Yes | 09/23 |
| (R)-9s |
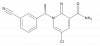 |
GPCR | ADRA1D | (S)-9s | Yes | 05/18 |
| BAY-6672 |
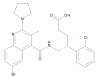 |
GPCR | PTGFR | BAY-403 | Yes | 11/20 |
| TP-020 |
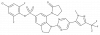 |
Transferase | MGAT2 | TP-020n | Yes | 11/18 |
| BAY-091 |
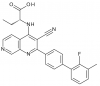 |
Kinase | PIP4K2A | BAY-0361 | No | 10/21 |
| BAY-826 |
 |
Kinase | TIE1, TEK, DDR1, DDR2 | BAY-309 | Yes | 05/18 |
| BAY-179 |
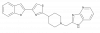 |
Complex I | BAY-070 | Yes | 01/20 | |
| IPP/CNRS-A017 |
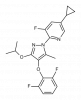 |
Dehydrogenase | DHODH | IPP/CNRS-A019 | No | 10/19 |
| WM-1119 |
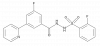 |
Transferase | KAT6A, KAT6B | WM-2474 | Yes | 06/20 |
|
IOX1 (tool compound) |
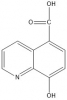 |
2OG | Pan-KDM | No | 01/12 | |
| BI-8668 |
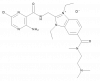 |
Sodium channels | Epithelial sodium channel (ENaC) | BI-0337 | Yes | 11/23 |
| A-079 |
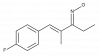 |
Ion channel | TRPA1 | A-226 | Yes | 11/18 |
| BI 605906 |
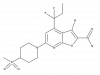 |
Kinase | IKBKB | BI-5026 | Yes | 06/21 |
| UCSF924 |
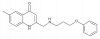 |
GPCR | DRD4 | UCSF924NC | Yes | 04/19 |
| BI-2081 |
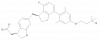 |
GPCR | FFAR1 | BI-0340 | Yes | 09/22 |
| PF-04418948 |
 |
GPCR | PTGER2 | PF-04475866 | Yes | 05/18 |
| BTZO-1 |
 |
Cytokine | MIF | BTZO-4 | No | 05/18 |
| BI-2540 |
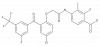 |
Reverse transcriptase | HIV NNRT | BI-2439 | Yes | 11/20 |
| WEB2086 |
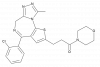 |
GPCR | PTAFR | WEB2387 | Yes | 01/24 |
| BAY-1797 |
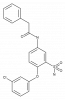 |
Ion channel | P2X4 | BAY-207 | Yes | 11/18 |
| PFI-653 |
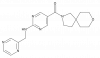 |
Hydrolase | VNN1 | PFI-653-N | Yes | 09/21 |
| ABT-724 |
 |
GPCR | DRD4 | A-769 | Yes | 05/18 |
| BAY-985 |
 |
Kinase | TBK1, IKBKE | BAY-440 | Yes | 01/20 |
| A-1596586 |
 |
Ion channel | CFTR | A-1596584 | No | 06/19 |
| BAY-7081 |
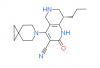 |
Phoshodiesterase | PDE9A | BAY-7424 | Yes | 09/23 |
| I-CBP112 |
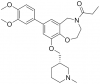 |
Bromodomains | CREBBP, EP300 | No | 03/13 | |
| BI-3802 |
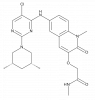 |
Transcription factor | BCL6 | BI-5273 | No | 11/23 |
| BAY-876 |
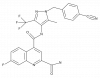 |
Solute carriers | SLC2A1 | BAY-588 | Yes | 05/18 |
| GSK620 |
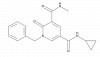 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | Yes | 02/21 | |
| BAY-293 |
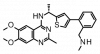 |
Nucleotide exchange factor | SOS1 | BAY-294 | No | 04/19 |
| TP-050 |
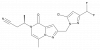 |
Ion channel | GRIN2A | TP-050n | Yes | 09/22 |
| GSM1 |
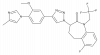 |
Protease | Gamma secretase complex | GSM-NC | Yes | 05/18 |
| NVS-MALT1 |
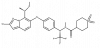 |
Protease | MALT1 | NVS-MALT1-C | Yes | 02/20 |
| JA397 |
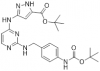 |
Kinase | CDK14-18 | JA314 | 07/23 | |
| JQ1 |
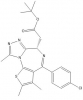 |
Bromodomains | BRD2, BRD3, BRD4, BRDT | (-)-JQ1 | Yes | 06/10 |
| PFI-2 |
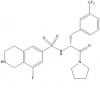 |
Methyltransferase | SETD7 | (S)-PFI-2 | No | 05/13 |
| LP99 |
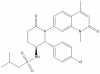 |
Bromodomains | BRD9, BRD7 | 2S,3R-enantiomer | No | 12/14 |
| A-196 |
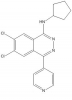 |
Methyltransferase | SUV420H1, SUV420H2 | A-197, SGC2043 | No | 07/15 |
| A-395 |
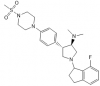 |
WD40 repeat | EED | A-395N | 07/16 | |
| BAY-299 |
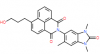 |
Bromodomains | TAF1, BRD1 | BAY-364 | No | 06/16 |
| BAY-598 |
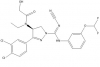 |
Methyltransferase | SMYD2 | BAY-369 | Yes | 04/15 |
| BAY-6035 |
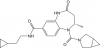 |
Methyltransferase | SMYD3 | BAY-444 | 06/17 | |
| BAY-850 |
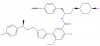 |
Bromodomains | ATAD2 | BAY-460 | 01/17 | |
| BAY-805 |
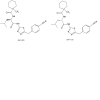 |
USP (ubiquitin specific protease) | USP21 | BAY-728 | No | 10/22 |
| SGC-CBP30 |
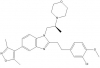 |
Bromodomains | CREBBP, EP300 | BDOIA513 | No | 03/13 |
| BI-9321 |
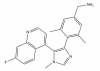 |
Methyl Lysine Binder | NSD3 | BI-9466 | 07/18 | |
| BI01383298 |
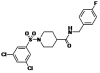 |
Solute carriers | SLC13A5 | BI01372674 | 01/18 | |
| CK156 |
 |
Kinase | STK17A, DRAK1 | CKJB71 | No | 10/21 |
| CS640 |
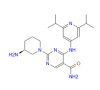 |
Kinase | CAMK1 | CS640s | Yes | 01/23 |
| L-Moses |
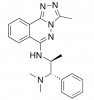 |
Bromodomains | KAT2A, KAT2B | D-Moses | Yes | 10/17 |
| DDR-TRK-1 |
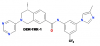 |
Kinase | DDR, TRK | DDR-TRK-1N | Yes | 08/18 |
| FM-381 |
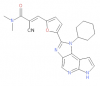 |
Kinase | JAK3 | FM-479 | 02/17 | |
|
GSK-J1 use pro-drug GSK-J4 for cell assays |
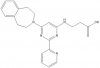 |
Lysine Demethylase | KDM6B/JMJD3, KDM6A/UTX, KDM5B/JARID1B | GSK-J2 (GSK-J5) | No | 07/12 |
| GSK484 |
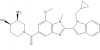 |
Arginine deiminases | PAD4 | GSK106 | Yes | 03/15 |
| GSK4027 |
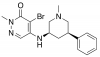 |
Bromodomains | KAT2A, KAT2B | GSK4028 | 06/17 | |
| GSK2801 |
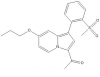 |
Bromodomains | BAZ2A, BAZ2B | GSK8573 | Yes | 12/14 |
| GSK8814 |
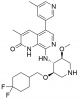 |
Bromodomains | ATAD2, ATAD2B | GSK8815 | 01/17 | |
| GSK6853 |
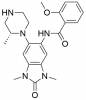 |
Bromodomains | BRPF1 | GSK9311 | Yes | 01/17 |
| GSK864 |
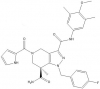 |
Dehydrogenase | IDH1 | GSK990 | Yes | 11/15 |
| JA310 |
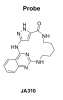 |
Kinase | MST3, MST4 | JA262 | 11/23 | |
| LLY-283 |
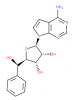 |
Methyltransferase | PRMT5 | LLY-284 | 07/16 | |
| M4K2234 |
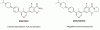 |
Kinase | ALK1, ALK2 | M4K2234NC | Yes | 02/22 |
| MRIA9 |
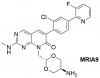 |
Kinase | SIK1, SIK2, SIK3 | MR7 | Yes | 08/21 |
| MRK-740 |
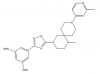 |
Methyltransferase | PRDM9 | MRK-740-NC | 06/18 | |
| MRK-952 |
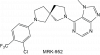 |
Hydrolase | NUDT5 | MRK-952-NC | No | 10/22 |
| MRK-990 |
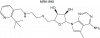 |
Methyltransferase | PRMT9, PRMT5 | MRK-990-NC | No | 10/22 |
| MS049 |
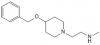 |
Methyltransferase | CARM1, PRMT6 | MS049N | No | 06/15 |
| MS023 |
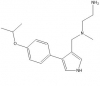 |
Methyltransferase | PRMT1, PRMT3, CARM1, PRMT6, PRMT8 | MS094 | No | 07/15 |
| MSC1186 |
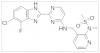 |
Kinase | SRPK | MSC5360 | 01/23 | |
| MU1210 |
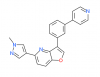 |
Kinase | CLK1, CLK2, CLK4, DYRK1A, DYRK1B, DYRK2 | MU140 | No | 05/19 |
| MU1700 |
 |
Kinase | ALK1, ALK2 | MU1700NC | Yes | 02/22 |
| MU1742 |
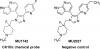 |
Kinase | CK1δ, CK1ε | MU2027 | Yes | 10/22 |
| Homer |
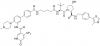 |
WD40 repeat | WDR5 | nc_WDR5 and nc_VHL | No | 07/21 |
| NR162 |
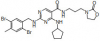 |
Kinase | CASK | NR187 | No | 09/20 |
| NVS-BPTF-1 |
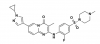 |
Bromodomains | BPTF | NVS-BPTF-C | No | 02/19 |
| NVS-CECR2-1 |
 |
Bromodomains | CECR2 | NVS-CECR2-C | No | 06/15 |
| NVS-MLLT-1 |
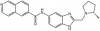 |
YEATS | MLLT1, MLLT3 | NVS-iMLLT-C | No | 03/20 |
| NVS-PAK1-1 |
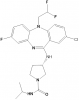 |
Kinase | PAK1 | NVS-PAK1-C | No | 06/16 |
| OICR-9429 |
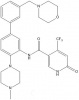 |
WD40 repeat | WDR5 | OICR-0547 | No | 05/14 |
| OICR11029 |
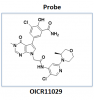 |
Transcription factor | BCL6 | OICR11600 | No | 01/24 |
| PFI-3 |
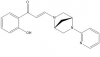 |
Bromodomains | SMARCA2, PBRM1, SMARCA4 | PFI-3oMet | No | 11/13 |
| PFI-5 |
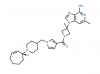 |
Methyltransferase | SMYD2 | PFI-5N | 07/17 | |
| PFI-6 |
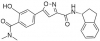 |
YEATS | MLLT1, MLLT3 | PFI-6N | No | 06/20 |
| PFI-7 |
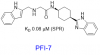 |
E3 ligase | GID4 | PFI-7N | No | 09/21 |
| PFI-8 |
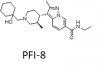 |
YEATS | YEATS | PFI-8N | No | 01/24 |
| SGC-AAK1-1 |
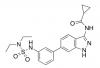 |
Kinase | AAK1, BMP2K, BIKE | SGC-AAK1-1N | 02/18 | |
| SGC-SMARCA-BRDVIII |
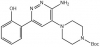 |
Bromodomains | PB1(5), SMARCA2/4 | SGC-BRDVIII-NC | 12/20 | |
| SGC-CAF382-1 |
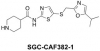 |
Kinase | CDKL5 | SGC-CAF268-1N | 09/23 | |
| SGC-CAMKK2-1 |
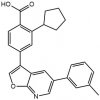 |
Kinase | CAMKK1, CAMKK2 | SGC-CAMKK2-1N | No | 01/20 |
| SGC-CDKL5_GSK3 |
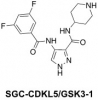 |
Kinase | CDKL5, GSK3 | SGC-CDKL5/GSK3-1N | No | 09/23 |
| SGC-GSK3-1 |
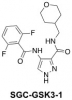 |
Kinase | GSK3 | SGC-CDKL5/GSK3-1N | No | 09/23 |
| SGC-CK2-1 |
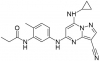 |
Kinase | CSNK2A1, CSNK2A2 | SGC-CK2-1N | No | 06/20 |
| SGC-CLK-1 |
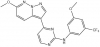 |
Kinase | CLK1, CLK2, CLK4 | SGC-CLK-1N | No | 02/20 |
| SGC-GAK-1 |
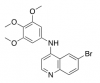 |
Kinase | GAK | SGC-GAK-1N | 02/18 | |
| SGC-PI5P4Kγ/MYLK4-1 |
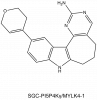 |
Kinase | PI5P4K gamma, MYLK4 | SGC-PI5P4Kγ/MYLK4-1N | No | 05/23 |
| SGC-PIKFYVE-1 |
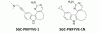 |
Kinase | PIKfyve | SGC-PIKFYVE-1N | No | 10/22 |
| SGC-STK17B-1 |
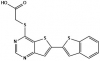 |
Kinase | STK17B, DRAK2 | SGC-STK17B-1N | 02/20 | |
| SGC-UBD1031 |
 |
UBD (ubiquitin binding domain) | SGC-UBD1031N | No | 05/24 | |
| SGC-UBD253 |
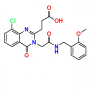 |
UBD (ubiquitin binding domain) | HDAC6 | SGC-UBD253N | No | 01/22 |
| SGC0946 |
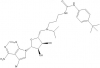 |
Methyltransferase | DOT1L | SGC0649 | No | 08/12 |
| GSK591 |
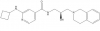 |
Methyltransferase | PRMT5 | SGC2096 | No | 07/15 |
| SGC3027 |
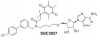 |
Methyltransferase | PRMT7 | SGC3027N | 03/18 | |
| SGC6870 |
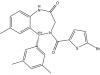 |
Methyltransferase | PRMT6 | SGC6870N | 06/19 | |
|
LLY-507 (Multiple Off-target Effects) |
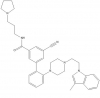 |
Methyltransferase | SMYD2 | SGC705 | No | 11/13 |
| SGC-CK2-2 |
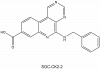 |
Kinase | CK2/CSNK2 | SGK-CK2-2N | 05/23 | |
| SKI-73 |
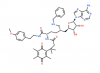 |
Methyltransferase | CARM1 | SKI-73N | 07/17 | |
| TH-257 |
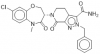 |
Kinase | LIMK1, LIMK2 | TH-263 | 07/23 | |
| TH-257 |
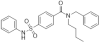 |
Kinase | LIMK1, LIMK2 | TH-263 | 07/17 | |
| TP-064 |
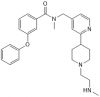 |
Methyltransferase | CARM1 | TP-064N | 06/16 | |
| TP-238 |
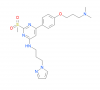 |
Bromodomains | CECR2, BPTF | TP-422 | 09/18 | |
| TP-472 |
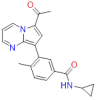 |
Bromodomains | BRD9, BRD7 | TP-472N | Yes | 06/16 |
| UNC0638 |
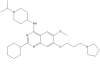 |
Methyltransferase | EHMT2/G9a, EHMT1/GLP | UNC0737 | No | 06/10 |
| UNC1215 |
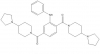 |
Methyl Lysine Binder | L3MBTL3 | UNC1079 | No | 08/12 |
| UNC1999 |
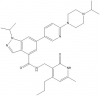 |
Methyltransferase | EZH2 | UNC2400 | Yes | 05/13 |
| UNC6934 |
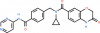 |
Methyl Lysine Binder | NSD2-PWWP1 | UNC7145 | 06/19 | |
| UNC8732 |
 |
Methyl Lysine Binder | NSD2 | UNC8884 | No | 01/24 |
| VinSpinIn |
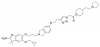 |
Spindlin domains | Spin1 | VinSpinIC | 09/18 | |
| SGC707 |
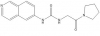 |
Methyltransferase | PRMT3 | XY1 | No | 06/14 |
Chemical Handles
This versatile modality can be used to develop tools for proximity-inducing chemistry, tracers for target engagement assays, PROTACs, chemical inducers of degradation, and pull-down reagents. Handles bind their target with Kd < 1 micromolar. Their selectivity and cell activity has not been determined.
| Chemical Handle | Chemical Probe | Structures | Protein family | Specific Targets |
Control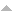 |
In Vivo Activity* | Added |
|---|---|---|---|---|---|---|---|
| PFI-E3H1 | PFI-7 |
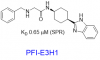 |
E3 ligase | GID4 | PFI-7N | No | 09/21 |
Please select a protein family
