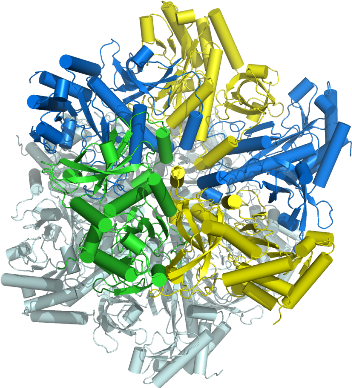
GS
PDB:2OJW
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:BC011852
Entry Clone Source:MGC
SGC Clone Accession:GLULA-s002
Tag:N-terminal hexahistidine tag with integrated TEV protease cleavagesite: mhhhhhhssgvdlgtenlyfq*sm
Host:BL21 (DE3) Gold pRare2
Construct
Prelude:Sequence:MHHHHHHSSGVDLGTENLYFQSMASSHLNKGIKQVYMSLPQGEKVQAMYIWIDGTGEGLRCKTRTLDSEPKCVEELPEWNFDGSSTLQSEGSNSDMYLVPAAMFRDPFRKDPNKLVLCEVFKYNRRPAETNLRHTCKRIMDMVSNQHPWFGMEQEYTLMGTDGHPFGWPSNGFPGPQGPYYCGVGADRAYGRDIVEAHYRACLYAGVKIAGTNAEVMPAQWEFQIGPCEGISMGDHLWVARFILHRVCEDFGVIATFDPKPIPGNWNGAGCHTNFSTKAMREENGLKYIEEAIEKLSKRHQYHIRAYDPKGGLDNARRLTGFHETSNINDFSAGVANRSASIRIPRTVGQEKKGYFEDRRPSANCDPFSVTEALIRTCLLNETG
Vector:pNIC-Bsa4
Growth
Medium:TB
Antibiotics:Procedure:30 microL BL21(DE3) gold pRARE2 cells were transformed with 2 microL plasmid miniprep for 30 min on ice followed by heatshock at 42degC for 45 sec. SOC, 125 microL, was added to the cellsuspension which was then incubated for 1 hour at 37degC and plated on LB-plates containing kanamycin (50 microG/mL). 30 mL TB (supplemented with 8 g/L glycerole, 100 µg kanamycin/ml and 3.4 microG/ml chloramphenicol) was inoculated with cells and grown overnight (ON) at 30degC. 20 ml of the inoculation culture was added to 1.5 L TB (supplemented 8g glycerole/L and 50 microG kanamycin/mL) in 2 L bottles. The flask was incubated in the LEX system-water bath at 37degC until OD600 reached 2. At this time the flask was transferred to an 18degC water bath in the LEX-system. Expression of protein was induced after approximately 1 hour by addition of 0.5 mM IPTG and continued for approximately 20 hours. Cells were harvested by centrifugation in a SLC-6000 rotor for 10 minutes at 5000 rpm (WCW 28.8 g). Pellets were suspended in 140 mL 50 mM Naphosphate pH 7.5, 10 % glycerol, 0.5 mM TCEP and 500 mM NaCl, 10mM imidazole and Complete EDTA-free protease inhibitor (Roche Biosciences). Suspended cells were stored at -80degC until further use.
Purification
ProcedureThe cleared lysate was loaded onto a HiTrap IMAC column (Amersham Biosciences) using an ÄktaExpress system. Eluted protein was run through a Superdex 200 16/60 gel filtration column. Fractions containing protein were pooled and the TCEP-concentration adjusted to 2 mM. Purified GS only remained stable in the presence of ATP and MnCl2 and thus 10 mM ATP and MnCl2 were added to the protein before concentration to 20 mg/ml.
Extraction
ProcedureBefore lyses, 8 microL of 250U/microL benzonase (Novagen) was added to the suspended cells and the sample was sonicated (Sonics VibraCell) at 80% amplitude for 3 min (pulse: 4 sec on and 4 sec off).The sample was spun for 30 min at 20500 rpm in a Sorvall SA-800 rotor. The soluble fraction was decanted and filtered through a 0.45 microM syringe filter
Concentration:LigandMassSpec:Crystallization:Crystals of GS were grown using vapor diffusion at 4degC by mixing equal amounts of protein solution at 20 mg/ml and reservoir solution containing 10% Isopropanol, 200 mM Sodium Chloride, 100 mM HEPES pH 7.5. Plate-like crystals appeared after one to three days.
NMR Spectroscopy:Data Collection:Data to 2.05 Å resolution were collected from a single crystal at ESRF (ID14-2), Grenoble, France. Crystal belonged to C2 space group with cell parameters of a=177.6 Å, b=122.6 Å, c=126.6 Å, α= 90°, β=130.6°, γ= 90°.
Data Processing:The structure was solved by molecular replacement using Glutamine synthetase from maize as a search model (PDB entry: 2D3A) with the program MolRep. The asymmetric unit contains a pentamer. The space group was C2 with cell dimensions a=177.6Å, b=122.6Å, c=126.6Å and beta=130.6. Refmac5 was used for refinement and Coot for model building. TLS restrained refinement was used in the refinement process. The TLS groups were selected using the tlsmd server
http://skuld.bmsc.washington.edu/~tlsmd/. Data in the interval 39.25-2.05Å resolution was used and at the end of the refinement the values for R= 16.2% and Rfree= 21.2%. A few residues are disordered and not visible in the electron density map. Coordinates for the crystal structure were deposited in the Protein Data Bank, accession code 2OJW.