|
Structure
|
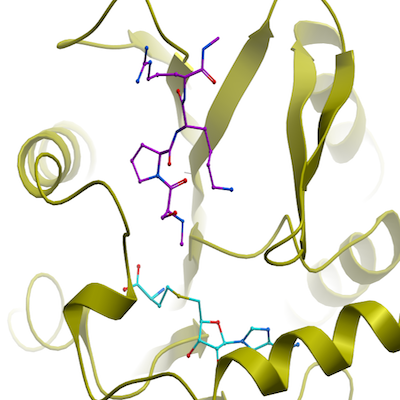
Crystal structure of human alpha N-terminal protein methyltransferase 1B
|
|
PDB Code
|
5UBB, 6DUB
|
|
Entry clone accession
|
NP_001129579
|
|
Entry clone source
|
|
|
SGC clone accession
|
JMC119-E02
|
|
Tag
|
N-terminal tag: mkkhhhhhhhhssglvprgsmsdsevnqeakpevkpevkpethinlkvsdgsseiffkikkttplrrlmeafakrqgkemdslrflydgiriqadqtpedldmedndiieahreqiggtenlyfqg
|
|
Construct sequence
|
Mkkhhhhhhhhssglvprgsmsdsevnqeakpevkpevkpethinlkvsdgsseiffkikkttplrrlmeafakrqgkemdslrflydgiriqadqtpedldmedndiieahreqiggtenlyfqg
TSQVINGEMQFYARAKLFYQEVPATEEGMMGNFIELSSPDIQASQKFLRKFVG
GPGRAGTDCALDCGSGIGRVSKHVLLPVFNSVELVDMMESFLLEAQNYLQVKG
DKVESYHCYSLQEFTPPFRRYDVIWIQWVSGHLTDKDLLAFLSRCRDGLKENGIII
LKDNVAREGCILDLSDSSVTRDMDILRSLIRKSGLVVLGQEKQDGFPEQCIPVWMFALH
|
|
Vector
|
pET28-MKH8SUMO
|
|
Expression host
|
BL21 (DE3) Codon plus RIL (Stratagene)
|
|
Growth method
|
NTMT2 was expressed in E.coli BL21 (DE3) codon plus RIL in Terrific Broth (TB) in the presence of 50 µg/mL of kanamycin. Cell were grown at 37 ºC to an OD600 of 1.5 and induced by isopropyl-1-thio-D-galactopyranoside (IPTG), final concentration 0.2 mM, and incubated overnight at 16 ºC. Cell pellets collected by centrifugation and frozen at -80 ºC.
|
|
Extraction buffers
|
Lysis buffer: 20 mM Tris-HCl pH 7.5, 400 mM NaCl, 5% glycerol and 2 mM beta-mercaptoethanol
|
|
Extraction procedure
|
Frozen cell pellet was thawed and suspended in lysis buffer. The cells were lysed by sonication (Virtis408912, Virsonic) on ice: the sonication protocol was 5 sec pulse at half-maximal frequency (5.0), 7 second rest, for 10 minutes total sonication time per pellet. The lysate was centrifuged at 15000rpm for 1h.
|
|
Purification buffers
|
Wash buffer: 20 mM Tris pH 7.5, 400 mM NaCl, 5% glycerol and 25 mM imidazole;
Elution buffer: 20 mM Tris pH 7.5, 400 mM NaCl, 5% glycerol and 300 mM imidazole;
Gel filtration buffer: 20 mM Tris-HCl pH 7.5, 150 mM NaCl and 0.5 mM TCEP
|
|
Purification procedure
|
Cells were lysed in 20 mM Tris-HCl pH 7.5, 400 mM NaCl, 5% glycerol and 2 mM beta-mercaptoethanol buffer and purified by Ni-NTA agarose chromatography. The SUMO tag was cleaved by TEV protease at 4 °C overnight and removed by reloading onto the Ni-NTA. The protein was diluted and applied onto HiTrap Q HP anion exchange chromatography column (GE Healthcare) equilibrated with 20 mM Tris-HCl pH 7.5, 25mM NaCl and 0.5 mM TCEP (tris (2-carboxyethyl) phosphine). The proteins were eluted with a linear gradient of 0-50% elution buffer (20 mM Tris-HCl pH 7.5, 1M NaCl and 0.5 mM TCEP). The proteins were further purified by gel filtration Superdex 200 10/300 (GE Healthcare). The gel filtration buffer contains 20 mM Tris-HCl pH 7.5, 150 mM NaCl and 0.5 mM TCEP.
|
|
Protein stock concentration
|
The purified protein was concentrated to 20 mg mL-1 using 15 mL concentrators with a 10,000 molecular weight cut-off (Amicon Ultra-15, UFC900524, Millipore).
|
|
Crystallization
|
NTMT2 purified from Escherichia coli contains endogenous methyl donor SAM/SAH, so we did not add any additional SAM/SAH during the crystallization. The NTMT2-SAM complex was crystallized in 20% (w/v) PEG3350 and 0.2 M sodium acetate via sitting drop vapor diffusion by mixing 1 microL protein and 1 microL reservoir solution at 4℃. To get the ternary complex crystal, the protein was incubated with SPKRIA peptide (from 100 mM stock) at a molar ratio of 1:1.5 for 1 h on ice before setting up the crystallization trial. The crystals of NTMT2 in complex with SPKRIA were obtained in 30% PEG2000 (w/v) and 0.1 M potassium thiocyanate at 18 ℃. The crystals were cryo-protected in the reservoir solution supplemented with 20% (v/v) glycerol and flash-frozen in liquid nitrogen.
|