The probe is available from Sigma and Cayman Chemical.
| Probe |
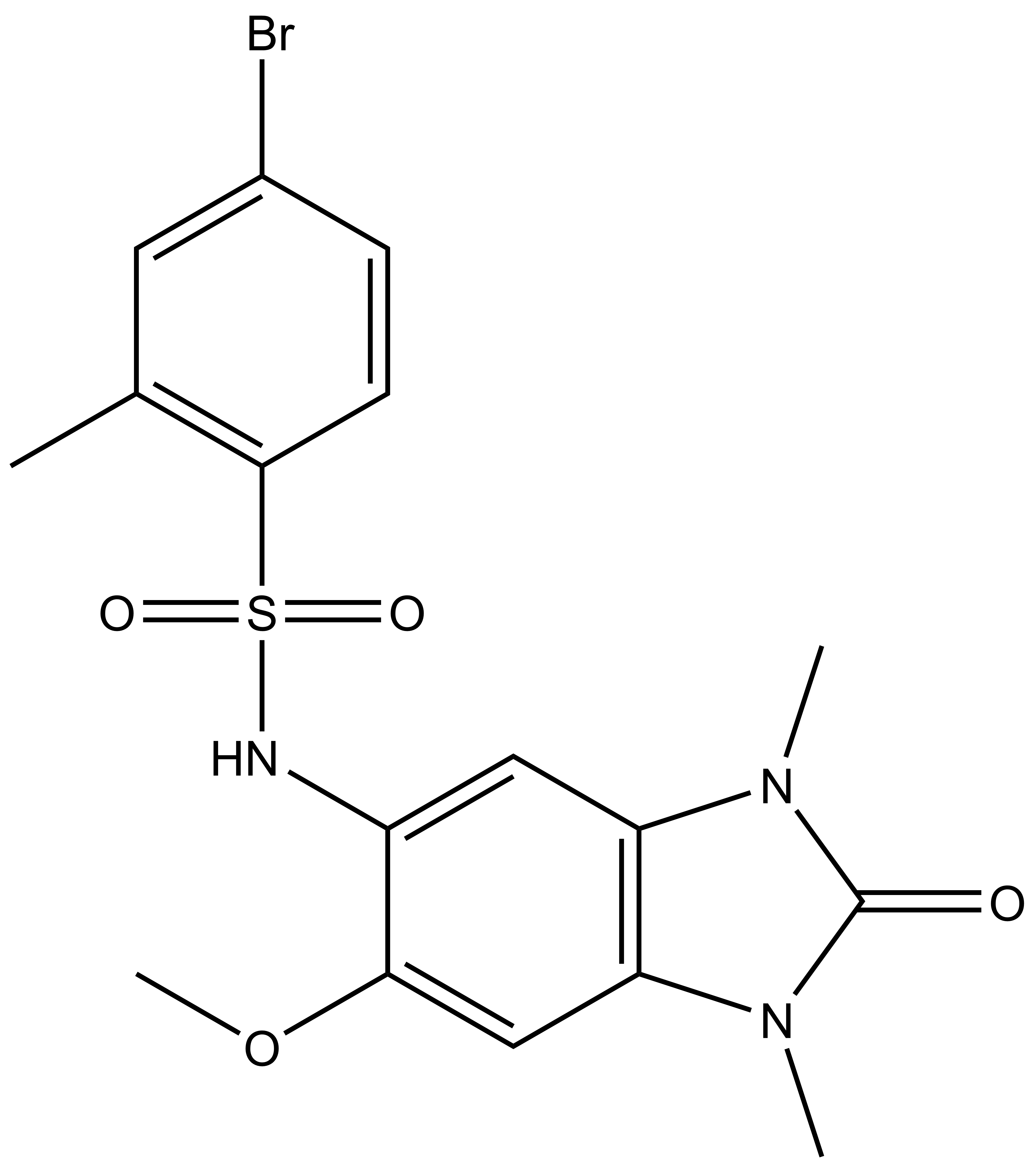 |
OF-1 |
A chemical probe for the bromodomains of the BRPF (BRomodomain and PHD Finger containing) family of proteins (BRPF1/2/3) has been discovered by the SGC.
BRPF1, BRPF2 (BRD1) and BRPF3 are scaffolding proteins, assembling HAT complexes of the MOZ/MORF family (MOZ, Ybf2/Sas3, Sas2, and Tip60) (1). These MYST complexes have a tetrameric core containing BRPF, the tumour suppressor ING and Eaf6/EPC (enhancer of polycomb)-related scaffold subunits. MYST complexes play crucial roles in DNA repair, recombination, and replication as well as in transcription activation (2,3). MOZ is frequently translocated in AML (acute myeloid leukemia) and is required for HSC proliferation (4).
Two BRPF1 isoforms (isoform A and B) can be generated by alternative splicing. In contrast to BRPF1B, the isoform A harbours a residue insertion in the ZA-loop that prevents binding to acetylated histone peptides (5).
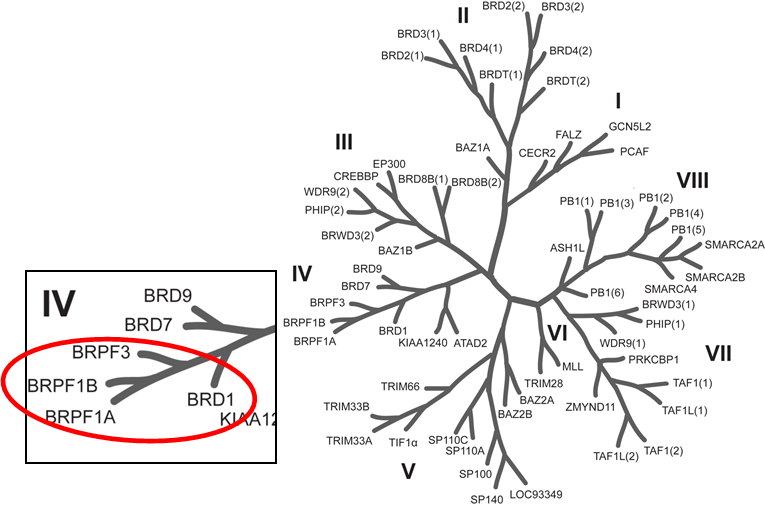
Phylogenetic tree of bromodomains and detailed view at the BRPF family.
OF-1 has been shown to bind to BRPF1B with a KD of 100 nM (ITC), to BRPF2 with a KD of 500 nM (ITC) and to BRPF3 with a KD of 2.4 µM (ITC). Selectivity against other BRDs is very good, in general >100-fold. The closest off-target effects are against BRD4 (39-fold selectivity) and TIF1α (50% inhibition at 20 µM in the alpha screen). OF-1 increases thermal stability in the cellular thermal shift assay (CETSA) of full length BRPF1B at 1 µM and also demonstrates accelerated FRAP recovery at 5 µM in the BRPF2 FRAP assay. It shows modest general cytotoxicity.
| Bromodomain | Kd/nM (ITC) | IC50/nM (Alpha Screen) | TM Shift °C |
|---|---|---|---|
| BRPF1B | 100 | 270 | 6.5±0.4 |
| BRPF2 (BRD1) | 500 | 2200 | 3.1±0.3 |
| BRPF3 | 2400 | ND | 3.3±0.2 |
| BRD4 (1) | 4000 | >10,000 | 2.0±0.2 |
OF-1 is a chemical probe with potent binding affinity for BRPF1 (KD of 100nM), BRPF2 (KD of 500 nM) and BRPF3 (KD of 2400 nM) as determined by isothermal titration calorimetry (ITC). Alpha screen biochemical assay confirmed OF-1 as a potent inhibitor of BRPF1 (IC50 of 270 nM).
OF-1 induced significant ΔTm shifts within the BRPF family. Weak interactions (2.1 °C) were also observed for BRD4(1), however alpha screen did not reveal strong interactions of OF-1 with BRD4 (IC50 of >10000 nM) and a KD of 4000nM for BRD4(1) reveals 39-fold selectivity when compared to BRPF1B isoform.
OF-1 shows no significant inhibition of protein kinases (<20% inhibition at 10 μM for 40 kinases screened).
We recommend to use the BRPF inhibitor NI-57 in parallel to confirm the result. Please do not apply OF-1 higher than 5 μM to avoid effects on BRD4 and verify the result by using NI-57.
OF-1 increases thermal stability in the CETSA of full length BRPF1B at 1 µM.
In a NanoBRETTM assay, BRPF1B, but not BRPF1A isoform shows dose-dependent displacement from histone H3.3, with IC50 of 0.08 μM and the FRAP assay reveals strong inhibition of BRPF2 (BRD1) at 5 µM concentration of OF-1.
OF-1 attenuates RANKL/MCSF induced differentiation of primary human monocytes into osteoclasts.
| Probe |
 |
OF-1 |
| Physical and chemical properties | |
|---|---|
| Molecular weight | 440.31 |
| Molecular formula | C17H18BrN3O4S |
| IUPAC name | 4-bromo-N-(6-methoxy-1,3-dimethyl-2-oxo-2,3-dihydro-1H-benzo[d]imidazol-5-yl)-2-methylbenzenesulfonamide |
| clogP | 2.55 |
| PSA | 78.95 |
| No. of chiral centres | 0 |
| No. of rotatable bonds | 4 |
| No. of hydrogen bond acceptors | 4 |
| No. of hydrogen bond donors | 1 |
| Storage | Stable as solid in the dark at -20°C. NB making aliquots rather than freeze-thawing is recommended |
| Dissolution | Soluble in DMSO |
Temperature Shift Assay
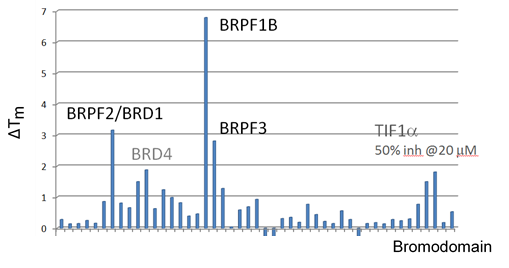
Selectivity screening of chemical probe OF-1 determined by temperature shift assay.
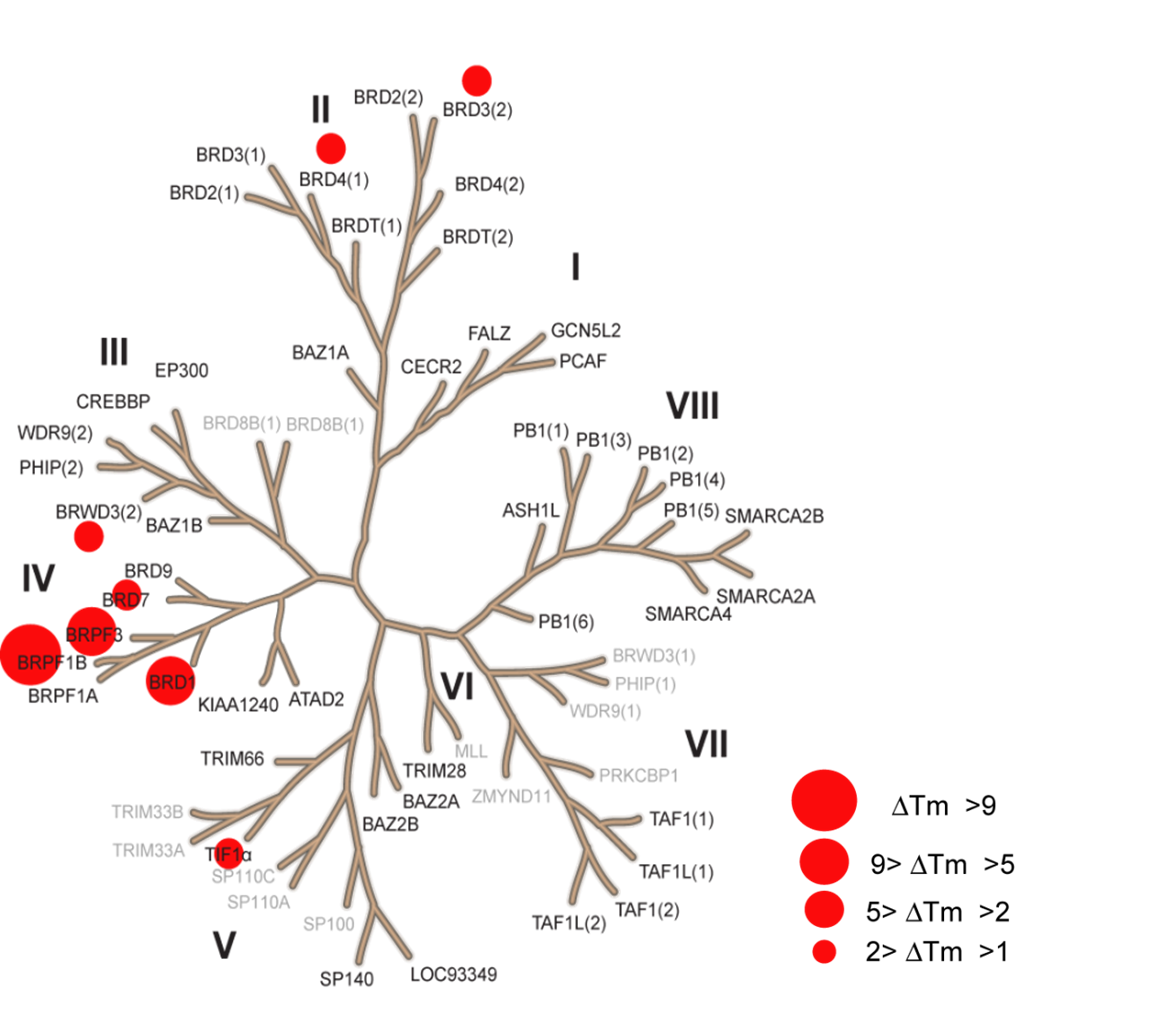

The temperature shifts mapped onto the phylogenetic tree using red circles corresponding to ΔTm as indicated in the figure.
CESTA
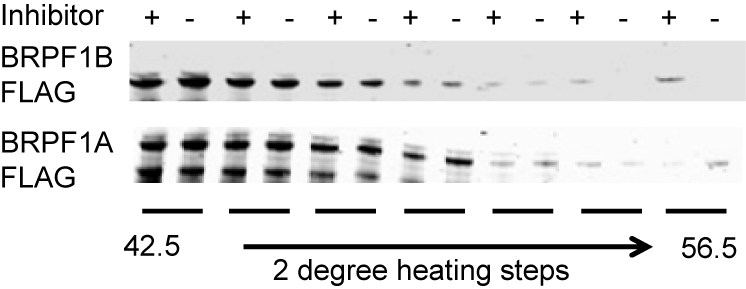
Thermal-stability of BRPF1A/B determined by CESTA at 1 µM OF-1.
NanoBRET
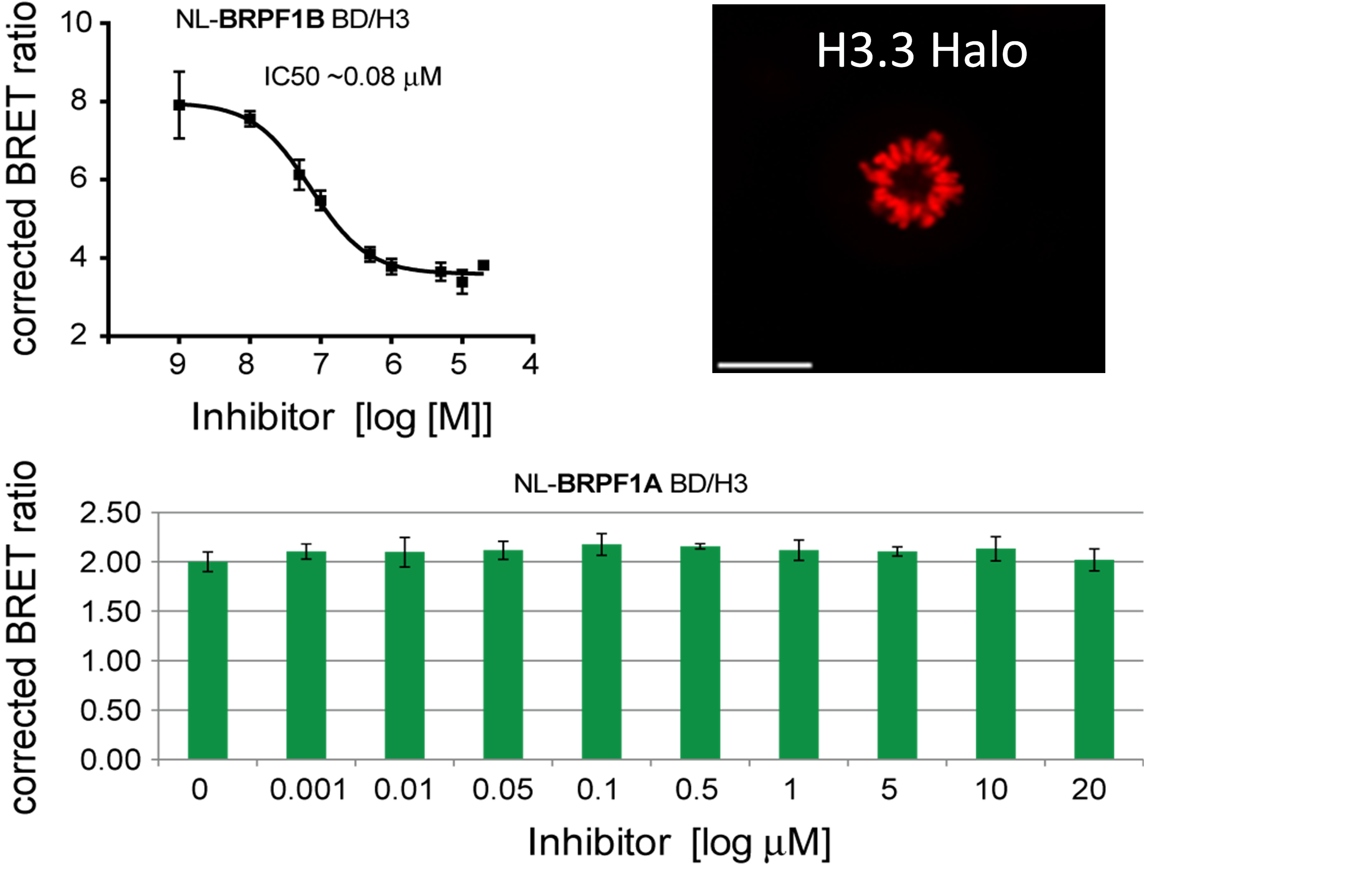
Dose-dependent displacement of BRPF1A/B from histone H3.3 following treatment with OF-1.
FRAP Assay
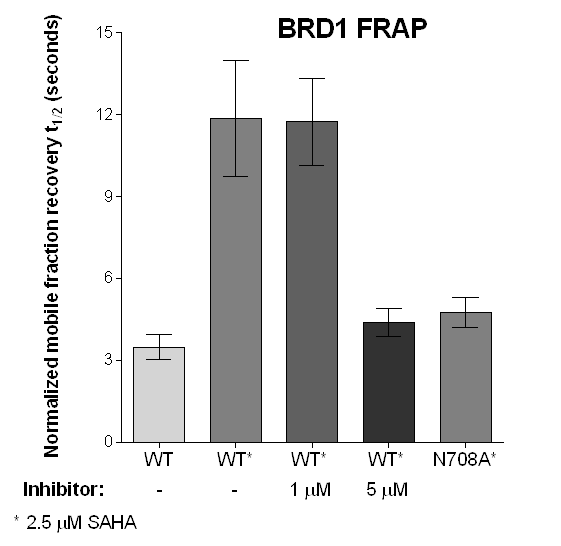
Half-times of fluorescence recovery (t1/2) after photo bleaching measured for BRPF2 (BRD1) after treatment with OF-1 at different concentrations with or without SAHA.
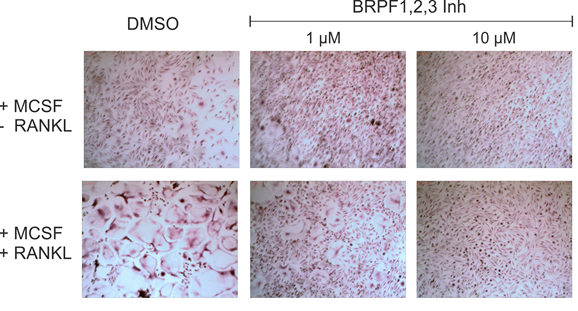
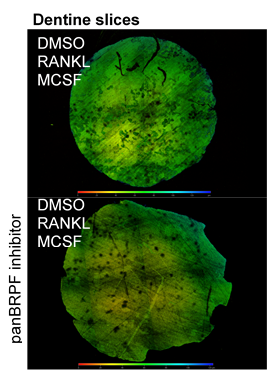
TRAP staining for osteoclasts and measurement of pits depth.
Work on this probe has been published in 'Selective targeting of Bromodomain-PHD fingers family (BRPF) bromodomains impairs osteoclast differentiation'.
Isothermal Titration Calorimetry (ITC)
All calorimetric experiments were performed on a VP-ITC micro-calorimeter (MicroCalTM, LLC Northampton, MA). Protein solutions were buffer exchanged by gel filtration or dialysis into buffer (20 mM Hepes pH 7.5, 150 mM NaCl, and 0.5 mM tris (2-carboxyethyl) phosphine (TCEP). All measurements were carried out at 288.15 K. All injections were performed using an initial injection of 2 µL followed by injections of 8 µL. The data were analysed with the MicroCal ORIGIN software package employing a single binding site model. The first data point was excluded from the analysis.
Temperature shift assay
Thermal melting experiments were carried out using a Stratagene Mx3005p Real Time PCR machine (Agilent Technologies). OF-1 was added at a final concentration of 10 µM. SYPRO Orange (Molecular Probes) was added as a fluorescence probe at a dilution of 1:1000 as described (6).
AlphaScreen Assay
Assays were performed as described previously with minor modifications (7). All reagents were diluted in 25 mM HEPES, 100 mM NaCl, 0.1 % BSA, pH 7.4 supplemented with 0.05 % CHAPS and allowed to equilibrate to room temperature prior to addition to plates. An 11-point 1:2.0 serial dilutions of the ligands was prepared on lowvolume 384-well plates (ProxiPlateTM-384 Plus, PerkinElmer, USA), using LabCyte Eco liquid handler. Plates filled with 5 µL of the assay buffer followed by 7 µL of biotinylated peptide [H-YSGRGKacGGKacGLGKacGGAKacRHRK(Biotin)-OH for BRD1, BRD4, BRPF1B and BRPF3 or YQTARKSTGGK(ac)APRKQLATKAK(biotin)-OH for TIF1α] and Histagged protein to achieve final assay concentrations of 25-100 nM depending on the dose-response curve for each individual protein. Plates were sealed and incubated for a further 30 minutes, before the addition of 8 µM of the mixture of streptavidin-coated donor beads (12.5 µg/mL) and nickel chelate acceptor beads (12.5 µg/mL) under low light conditions. Plates were foil-sealed to protect from light, incubated at room temperature for 60 minutes and read on a PHERAstar FS plate reader (BMG Labtech, Germany) using an AlphaScreen 680 excitation/570 emission filter set. IC50 values were calculated in Prism 5 (GraphPad Software, USA) after normalization against corresponding DMSO controls.
Fluorescence Recovery After Photobleaching (FRAP) Assay
FRAP studies were performed using U20S cells expressing full-length BRPF2 (BRD1). Six hours after transfection 2.5 µM SAHA (to increase global histone acetylation) was added and cells were treated with 1 µM or 5 µM of OF-1 1 hour before imaging and half recovery times from the fluorescence signal of the bleached U2OS nuclei were plotted.
NanoBRET
U2OS cells were co-transfected with Histone H3.3-HaloTag and NanoLuc-BRPF1. Twenty hours post-transfection cells were collected, washed with PBS, and exchanged into media containing phenol red-free DMEM and 4% FBS in the absence (control sample) or the presence (experimental sample) of 100 nM NanoBRET 618 fluorescent ligand (Promega). Cells were then treated with an increasing dose of OF-1. Five minutes prior to reading, NanoBRET furimazine substrate (Promega) was added to both control and experimental samples and plates were read on a CLARIOstar (BMG) equipped with 450/80 nm bandpass and 610 nm longpass filters with a 0.5 sec reading setting. A corrected BRET ratio was calculated and is defined as the ratio of the emission at 610 nm/450 nm for experimental samples (i.e. those treated with NanoBRET fluorescent ligand) subtracted by and the emission at 610 nm/450 nm for control samples (not treated with NanoBRET fluorescent ligand). BRET ratios are expressed as milliBRET units (mBU), where 1 mBU corresponds to the corrected BRET ratio multiplied by 1000. Relative IC50 values were estimated by non-linear regression analysis of (log) concentration of each inhibitor versus milliBRET ratios (GraphPad Prism).
Human osteoclast differentiation
Primary human peripheral blood mononuclear cells (PBMCs) were collected from a Histopaque generated buffy coat after gradient centrifugation at 20 min and 500g, brakes off. The CD14+ monocyte fraction was obtained by on-column CD14+-MACS bead isolation, washed twice with MACS buffer, and seeded at a density of 50 000 c/mL in αMEM/10%FCS supplemented with 25 ng/mL MCSF. After 6 days at 37 °C, 5% CO2 treatment with OF-1 with and without 50 ng/mL RANKL was started. Media were changed with fresh compound every 3−4 days. After 14−21 days, cells were fixed and stained for TRAP.
Bone Resorption Assays.
PBMCs were isolated and seeded onto self-cut dentine slices from ivory. After 14 days of differentiation, cells were removed from dentine slices. Dentine pits were imaged with confocal microscopy.