This probe is available from Tocris , Sigma and Cayman Chemical.
BI-9564 and a negative control are available from opnMe.com.
| Probe |
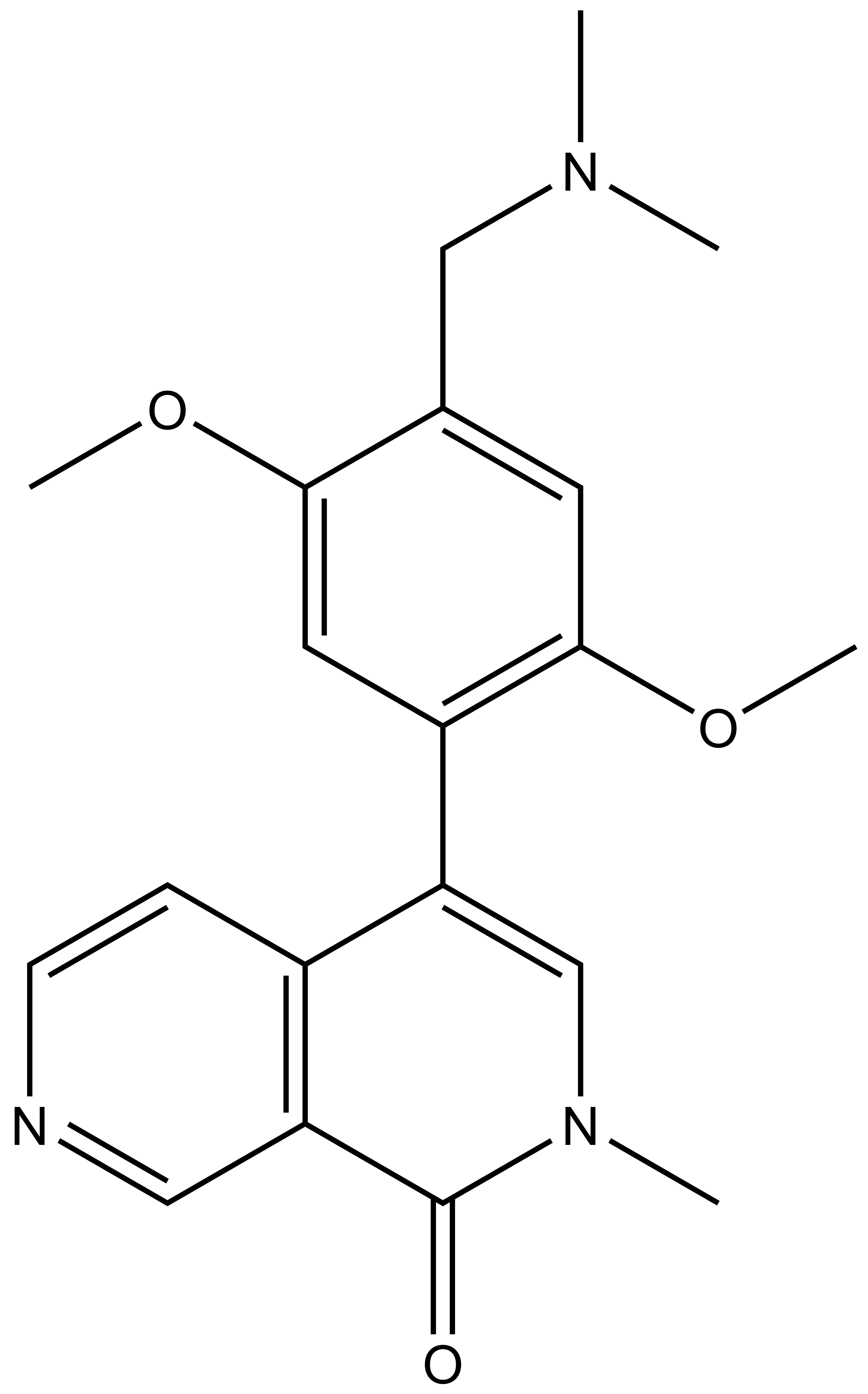 |
BI-9564 |
BRD7 and BRD9 are related bromodomain-containing proteins that form a small sub-branch of the bromodomain family tree (1). Human BRD9 contains a single bromodomain and has five isoforms that are produced by alternative splicing. Little is known about BRD9 function; BRD7 has been reported to act both as coactivator, for example for some p53 target genes and as corepressor, negatively regulating the GSK3B phosphotransferase activity. Both proteins have been implicated in chromatin remodelling as part of the SWI/SNF complex. BRD7 has been described as transcriptional corepressor that down-regulates the expression of target genes. Its binding to promoters also leads to increased histone H3 acetylation at 'Lys-9' (H3K9ac).
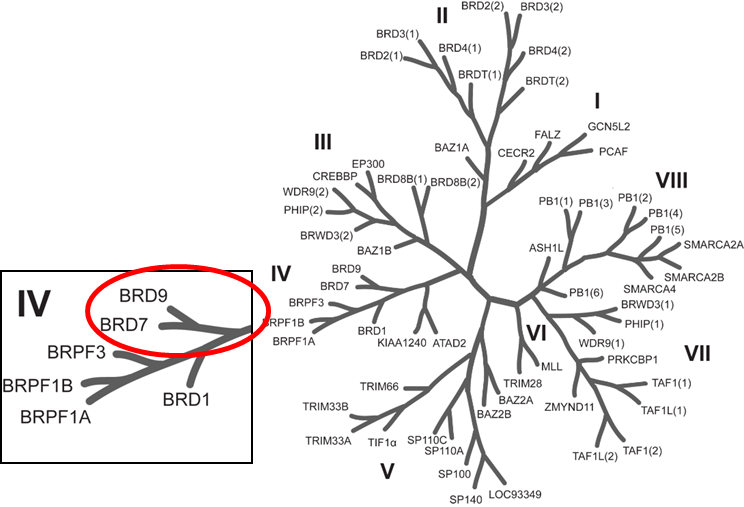
Phylogenetic tree of bromodomains and detailed view at the sub-branch of related bromodomain-containing proteins BRD7/9.
BI-9564 is a BRD9/7 specific inhibitor that has been developed in collaboration with Boehringer-Ingelheim. This probe was discovered through fragment-based screening and optimized by structure guided design.
BI-9564 binds to BRD9 with a higher potency (KD of 14 nM) than to BRD7 (KD of 239nM) as determined ITC, with CECR2 as the only off-target in vitro. BI-9564 is >30-fold selective against other non-Class IV bromodomains and negative against BET family members. BI-9564 does not significantly affect 324 kinases. BI-9564 is highly potent in cells and shows anti-proliferative activity in particular cell type.
BI-9564 binds to BRD9 with a higher affinity (KD of 14 nM, ITC) than to BRD7 (KD of 239nM, ITC), is completely negative on BET family members (>100 µM by alpha screen) and demonstrates cellular activity by FRAP on BRD9 and BRD7 at 0.1 µM and 1 µM, respectively.
| BRD | Kd/nM (ITC) | IC50/nM (Alpha Screen) |
| BRD7 | 239 | 3410 |
| BRD9 | 14 | 75 |
| CECR2 | 258 | NT |
Alpha screen confirmed BI-9564 as a potent inhibitor of BRD9 (IC50 of 75 nM). BI-9564 is very selective against other non-Class IV bromodomains, including the BETs. The only off-target effect was determined against CECR2 (18-fold selective), but not in cells (at 1 µM, FRAP). BI-9564 shows no significant off-target pharmacology against a panel of 324 kinases at concentrations less than 5 µM and is functionally active in cellular assays.
FRAP assay demonstrate that BI-9564 is able to disrupt the binding of BRD7 and BRD9 to chromatin in cells at 1 µM and 0.1 µM, respectively. BI-9564 shows no cellular inhibition of CECR2 at 1 µM.
In a broad cancer cell line panel treatment with BI-9564 resulted in selective growth inhibition of EOL-1 AML cells both in vitro (EC50 = 800 nM) and in a disseminated mouse model of AML (180 mg/kg/day).
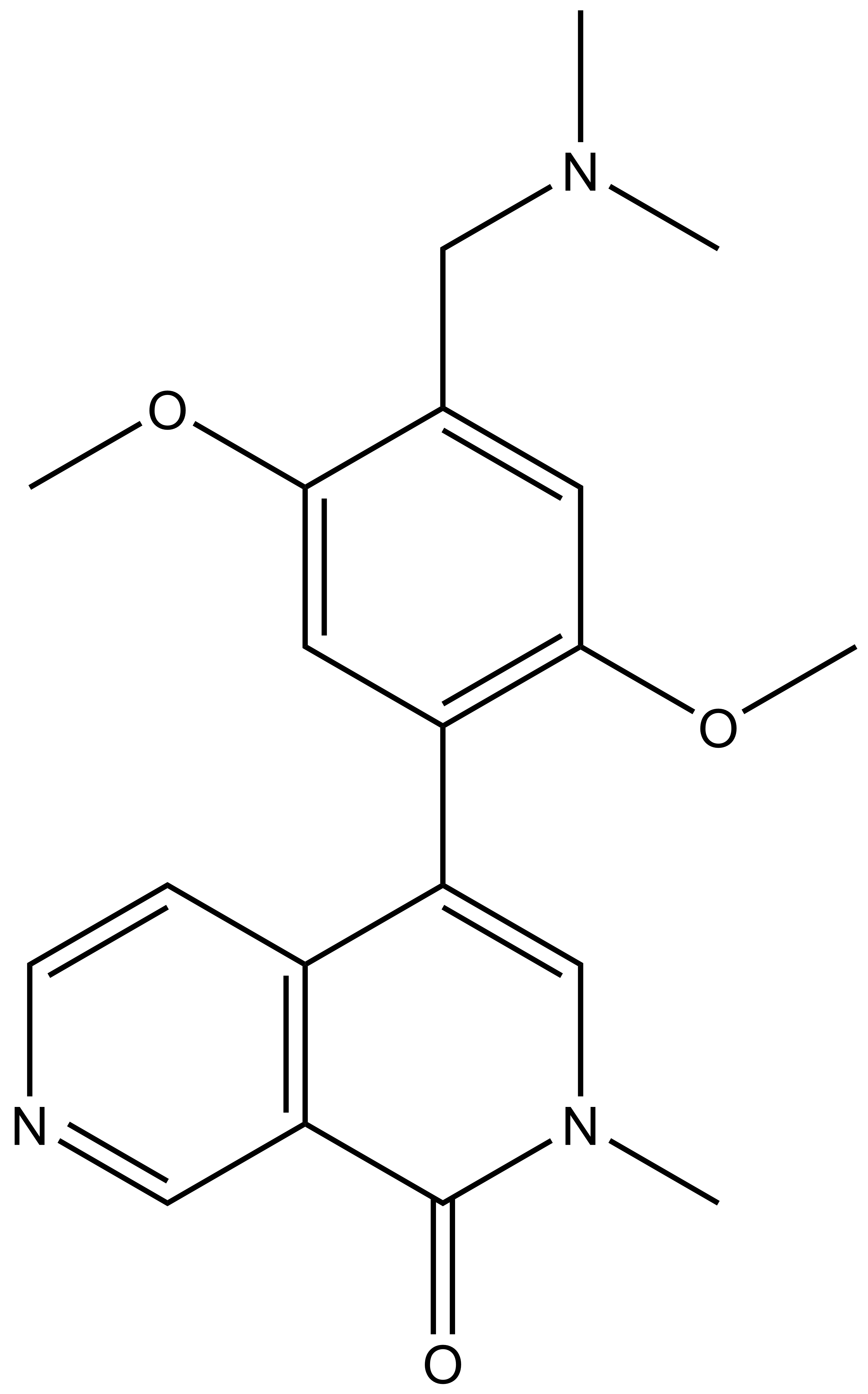 |
| 4-(4-((dimethylamino)methyl)-2,5-dimethoxyphenyl)-2-methyl-2,7-naphthyridin-1(2H)-one |
| Physical and chemical properties | |
|---|---|
| Molecular weight | 353.422 |
| Molecular formula | C20H23N3O3 |
| IUPAC name | 4-(4-((dimethylamino)methyl)-2,5-dimethoxyphenyl)-2-methyl-2,7-naphthyridin-1(2H)-one |
| clogP | 1.5 |
| PSA | 54.9 |
| No. of chiral centres | 0 |
| No. of rotatable bonds | 5 |
| No. of hydrogen bond acceptors | 5 |
| No. of hydrogen bond donors | 0 |
| Storage | Stable as solid in the dark at -20°C. NB making aliquots rather than freeze-thawing is recommended |
| Dissolution | Soluble in DMSO |
Temperature Shift Assay
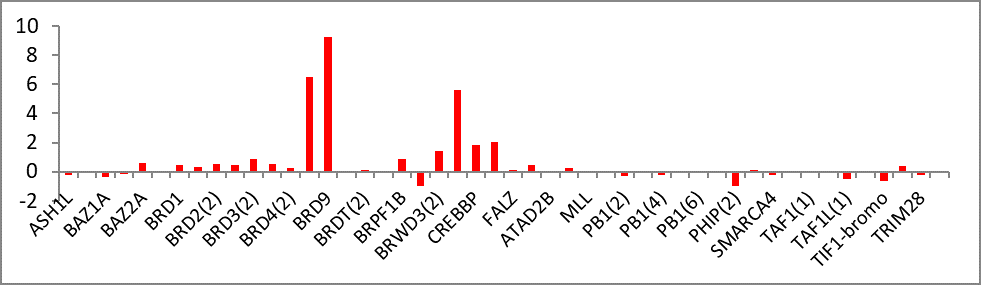
Selectivity profile of BI-9564 using temperature shift assay at 10µM.
Isothermal Titration Calorimetry (ITC)
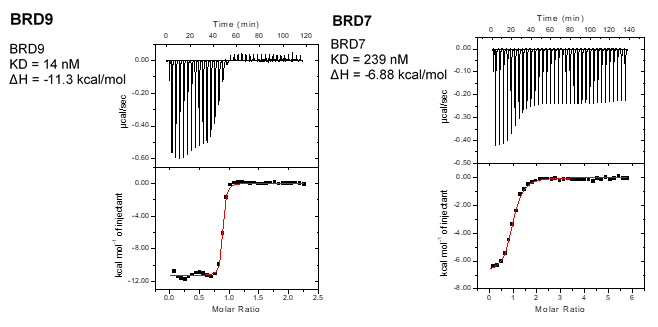
Binding affinity of BI-9564 towards BRD 7 and BRD 9 measured by ITC.
Fluorescence Recovery After Photobleaching (FRAP) Assay
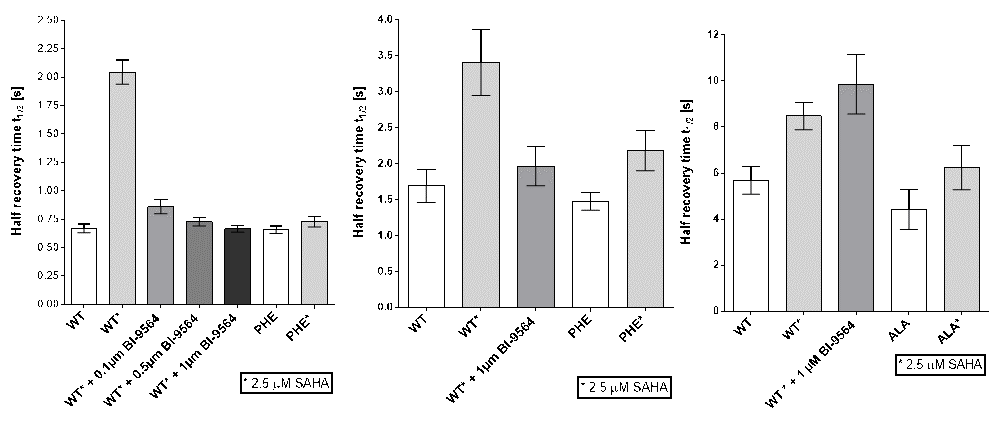
Half-times of fluorescence recovery (t1/2) after photo-bleaching measured for BRD9, BRD7 and CECR2 treated either with or without SAHA and BI-9564 at indicated concentrations.
Work on this probe has been published in ‘Structure-based design of an in vivo active selective BRD9 inhibitor’.
Isothermal Titration Calorimetry (ITC)
All calorimetric experiments were performed on a VP-ITC micro-calorimeter (MicroCalTM, LLC Northampton, MA). Protein solutions were buffer exchanged by gel filtration or dialysis into buffer (20 mM Hepes pH 7.5, 150 mM NaCl, and 0.5 mM tris (2-carboxyethyl) phosphine (TCEP). All measurements were carried out at 288.15 K. All injections were performed using an initial injection of 2 µL followed by injections of 8 µL. The data were analysed with the MicroCal ORIGIN software package employing a single binding site model. The first data point was excluded from the analysis.
Temperature shift assay
Thermal melting experiments were carried out using a Stratagene Mx3005p Real Time PCR machine (Agilent Technologies). BI-9564 was added at a final concentration of 10 µM. SYPRO Orange (Molecular Probes) was added as a fluorescence probe at a dilution of 1:1000 as described (2).
Fluorescence Recovery After Photobleaching (FRAP) Assay
U2OS cells were transfected (Fugene HD; Roche) with mammalian over- expression constructs encoding GFP fused to the N-terminus of full length BRD9, Brd7 or CECR2, respectively. Mutant proteins mutating the conserved Asn to Phe or Ala were generated as described in (3). The following mutations were introduced: N140A for human CECR2, N211F for mouse Brd7 and N163F for human BRD9. The imaging systemconsisted of a Zeiss LSM 710 laser-scanning and control system (Zeiss) coupled to an inverted Zeiss Axio Observer.Z1 microscope equipped with a high-numerical-aperture (N. A. 1.3) 40 x oil immersion objective (Zeiss). Samples were placed in an incubator chamber in order to maintaining temperature and humidity. FRAP and GFP fluorescence imaging were both carried out with an argon-ion laser (488 nm) and with a PMT detector set to detect fluorescence between 500-550 nm. Once an initial scan had been taken, a region of interest corresponding to approximately 50 % of the entire GFP positive nucleus was empirically selected for bleaching. A time lapse series was then taken to record GFP recovery using 1% of the power used for bleaching. The image datasets and fluorescence recovery data were exported from ZEN 2009, the microscope control software, into Origin to determine the average half-time for full recovery for 10-20 cells per treatment point.