| Probe | Negative control | |
 |
|  |
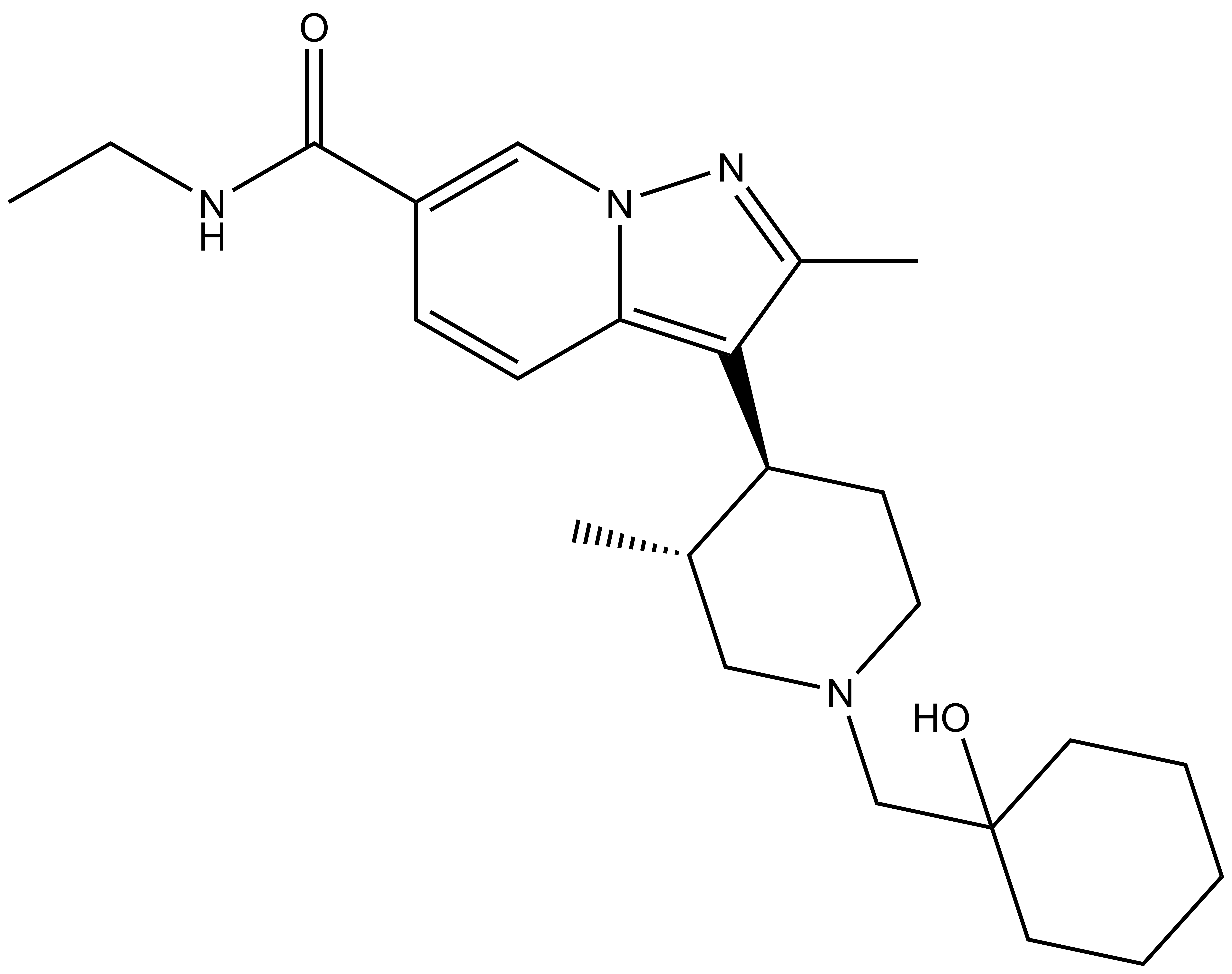
PFI-8 |
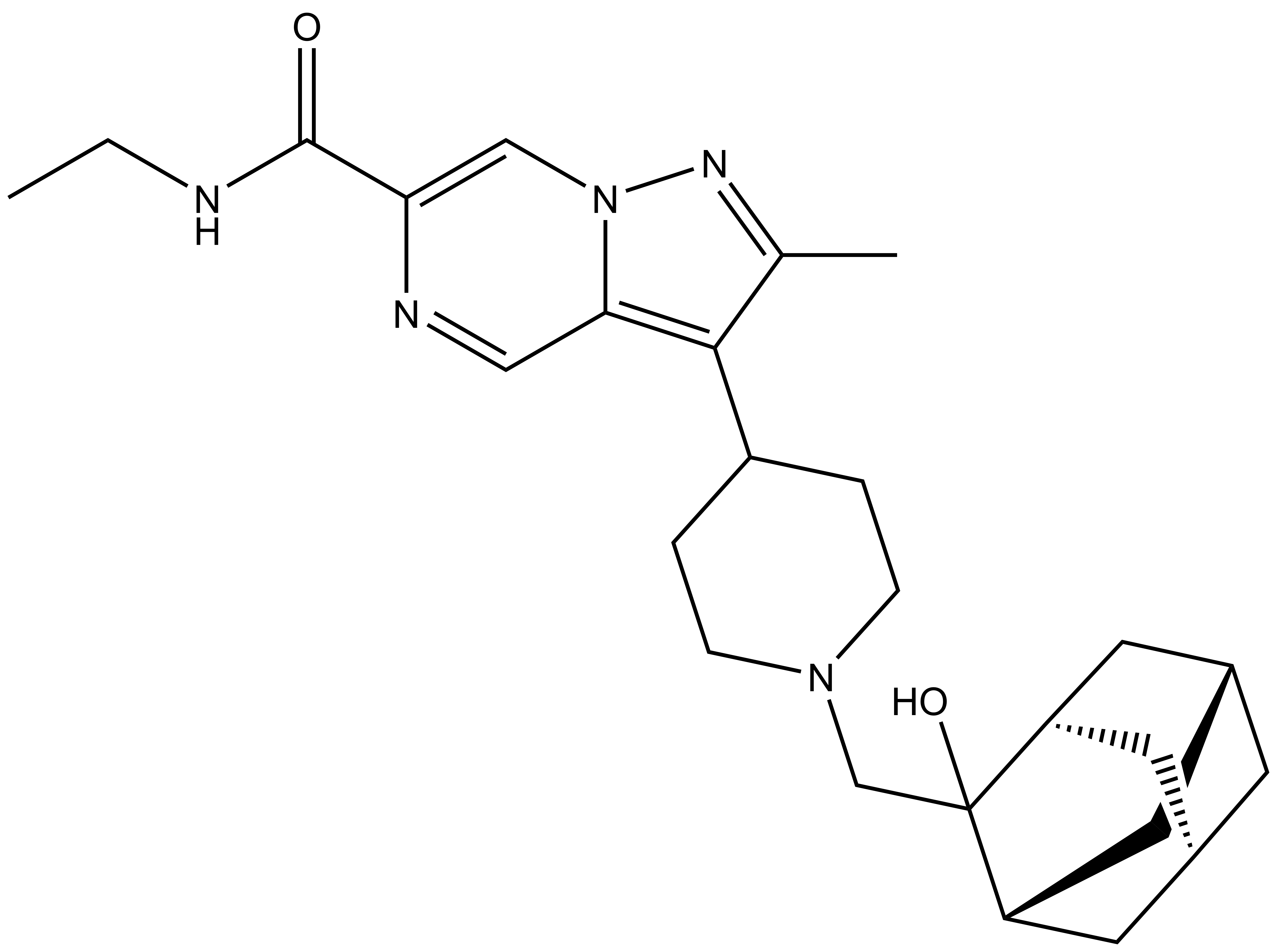
| PFI-8N |
The SGC in collaboration with Pfizer has discovered a potent chemical probe for the YEATS family, PFI-8. The YEATS-domain containing proteins are responsible for epigenetic signaling via acylated lysines including acetylation and higher order acylations such as propionylation, butyrylation, and crotonylation. PFI-8 binds YEATS 1-4 with FRET Kis of 495, 330, 462, and 33 nM respectively using a biotin-tagged crotonylated H3 peptide. PFI-8N, a negative control compound showed FRET Ki of >20 mM.
Binding of YEATS4 to PFI-8
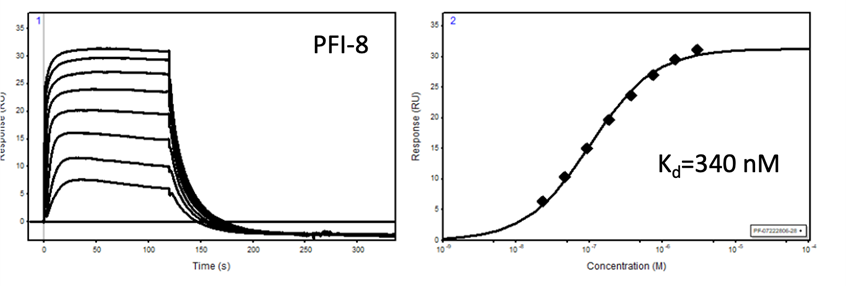
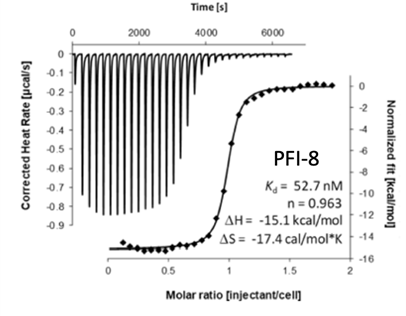
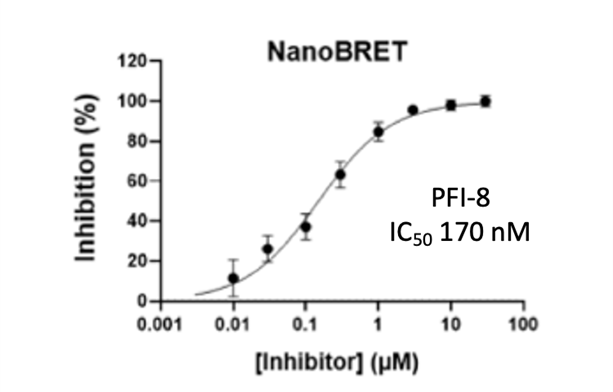
Cellular activity of PFI-8 using NanoBRET assay in HEK293 cells
1) (a) Heidenreich, D.; Moustakim, M.; Schmidt, J.; Merk, D.; Brennan, P. E.; Fedorov, O.; Chaikuad, A.; Knapp, S. Structure-based approach toward identification of inhibitory fragments for eleven-nineteen-leukemia protein (ENL). J. Med. Chem. 2018, 61, 10929−10934. (b) Moustakim, M.; Christott, T.; Monteiro, O. P.; Bennett, J.; Giroud, C.; Ward, J.; Rogers, C. M.; Smith, P.; Panagakou, I.; Diaz-Saez, L.; Felce, S. L.; Gamble, V.; Gileadi, C.; Halidi, N.; Heidenreich, D.; Chaikuad, A.; Knapp, S.; Huber, K. V. M.; Farnie, G. Heer, J.; Manevski, N.; Poda, G.; Al-Awar, R.; Dixon, D. J.; Brennan, P. E.; Fedorov, O. Discovery of an MLLT1/3 YEATS domain chemical probe. Angew. Chem., Int. Ed. 2018, 57, 16302−16307. (c) Christott, T.; Bennett, J.; Coxon, C.; Monteiro, O.; Giroud, C.;Beke, V.; Felce, S. L.; Gamble, V.; Gileadi, C.; Poda, G.; Al-Awar, R.;Farnie, G.; Fedorov, O. Discovery of a Selective Inhibitor for the YEATS Domains of ENL/AF9. SLAS Discov. 2019, 24, 133−141. (d) Ni, X.; Heidenreich, D.; Christott, T.; Bennett, J.; Moustakim, M.; Brennan, P. E., Fedorov, O.; Knapp, S.; Chaikuad, A. Structural insights into interaction mechanisms of alternative piperazine-urea YEATS domain binders in MLLT1. ACS Med. Chem. Lett. 2019, 10, 1661−1666. (e) Structural Genomics Consortium (SGC): PFI-6 A novel chemical probe for MLLT1/3. https://www.thesgc.org/chemicalprobes/PFI-6 (accessed June 23, 2020). (f) Structural Genomics Consortium (SGC): NVS-MLLT-1 A Potent and Selective inhibitor of YEATS proteins. https://www.thesgc.org/chemical-probes/NVS-MLLT-1 (accessed March 5, 2020) (g) Asiaban, J. N.; Milosevich, N.; Chen, E.; Bishop, T. R.; Wang, J.; Zhang, Y.; Ackerman, C. J.; Hampton, E. N.; Young, T. S.; Hull, M. V., Cravatt, B. F., Erb, M. A. Cell-based ligand discovery for the ENL YEATS domain. ACS Chem. Biol. 2020, 15, 895–903. (h) Ni, X.; Londregan, A. T.; Owen, D. R.; Knapp, S.; Chaikuad, A. ACS Chem. Bio. Structure and Inhibitor Binding Characterization of Oncogenic MLLT1 Mutants. 2021, 16, 569-576. (i) Listunov, D.; Linhares, B. M.; Kim, E.; Winkler, A.; Simes, M. L.; Weaver, S.; Cho, H. J.; Rizo, A.; Zolov, S.; Venkateshwar G. K.; Grembecka, J.; Cierpicki, T. Development of potent dimeric inhibitors of GAS41 YEATS domain. Cell Chem.Bio. 2021, 28, 1716-1727.
2) Measurement of ATP-depletion over 72 hours: Green, N.; Aleo, M. D.; Louise-May, S.; Price, D. A.; Will, Y. Using an in vitro cytotoxicity assay to aid in compound selection for in vivo safety studies. Bioorg. Med. Chem. Lett. 2010, 20, 5308-5312.