SGC707 is available from Cayman Chemical and Sigma and from Tocris
XY1 (the negative control of SGC 707) is available from Tocris and from Cayman Chemical.
| Probe | Negative control | |
 |
| 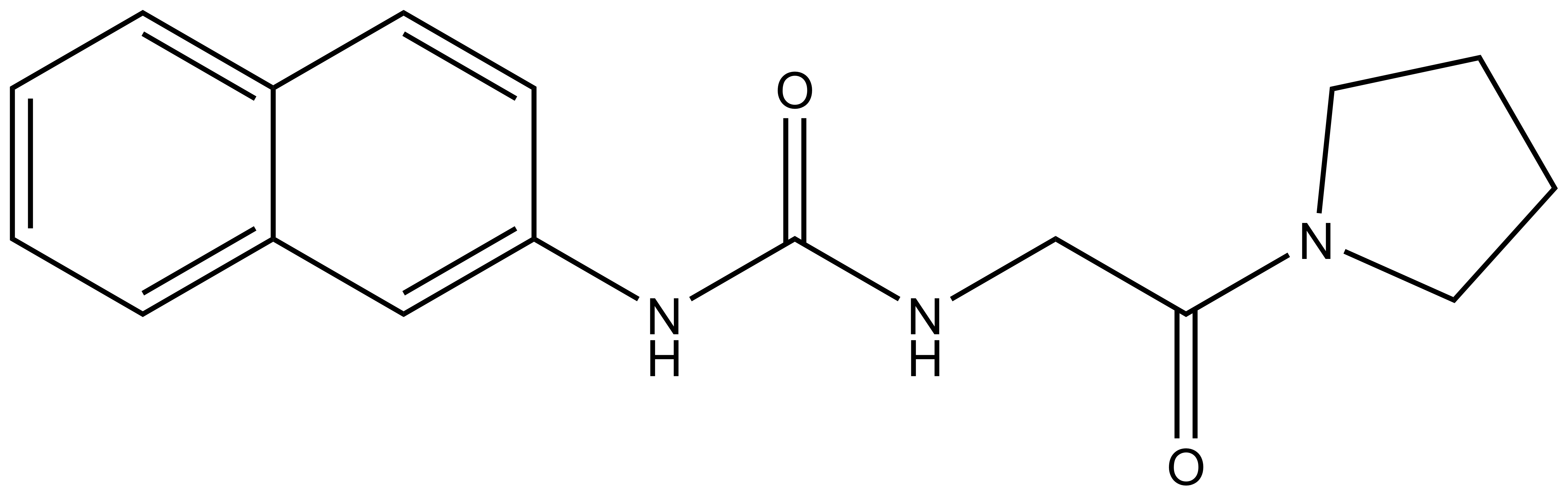 |
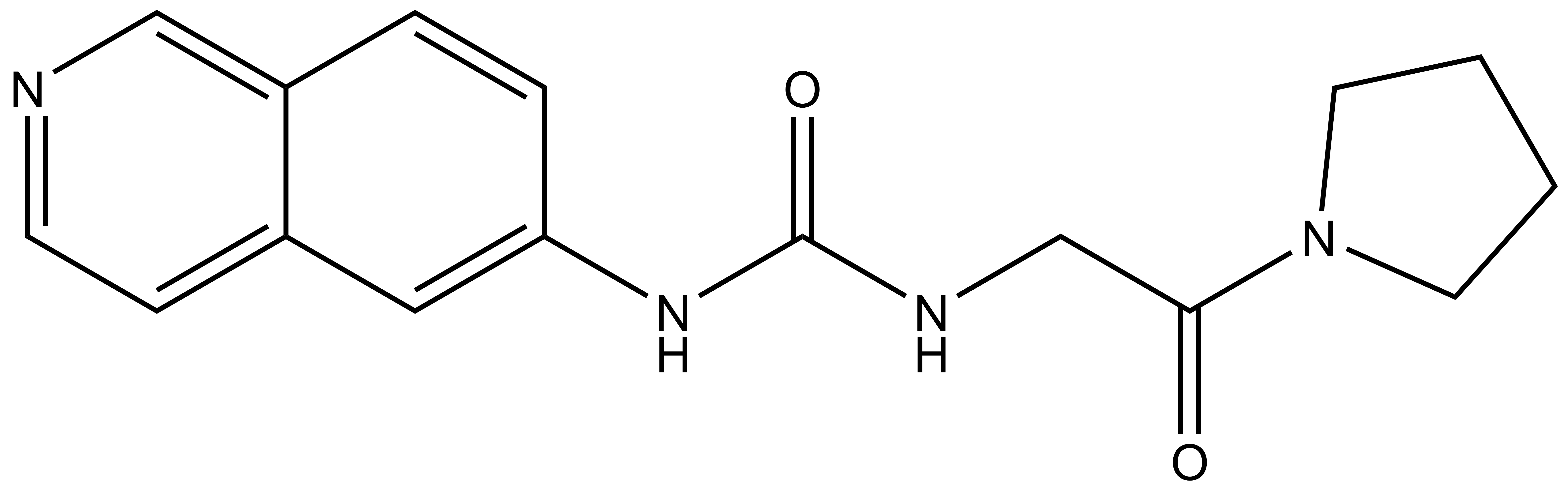
SGC707 (IC50=31 nM) |
| XY1 (IC50>100,000 nM) |
A collaboration between the SGC, China Novartis Institutes for Biomedical Research, and the center for Integrative Chemical Biology and Drug Discovery (CICBDD) at the University of North Carolina has resulted in the discovery of SGC707, a chemical probe for PRMT3, a protein which catalyzes the asymmetrical di-methylation of arginine residues. SGC707 is a potent allosteric inhibitor of PRMT3 (IC50 = 50 nM) with >100-fold selectivity over other methyltransferases, and other non-epigenetic targets. SGC707 has been shown to bind to PRMT3 with KD of 50 nM (ITC), and inhibits the methylation of histones in cells with IC50 value below 1 µM.
| Probe | Negative control | |
 |
|  |
SGC707 (IC50=31 nM) |
| XY1 (IC50>100,000 nM) |
Click here to download SDF file
| Physical and chemical properties | |
|---|---|
| Molecular weight | 298.1430 |
| Molecular formula | C16H18N4O2 |
| IUPAC name | 2-((3-aza-bicyclo[4.4.0]deca-1,3,5,7,9-pentaen-8-ylamino)-formylamino)-1-(pyrrolidin-1-yl)-ethanone |
| logP | 1.8 |
| PSA | 59.9 A |
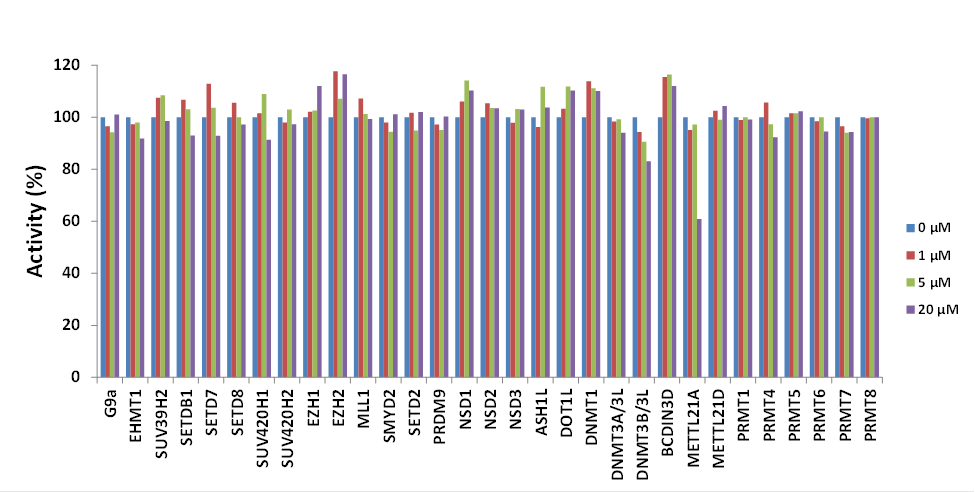
(C) Effect of SGC707 on the activity of 27 protein methyltransferases as well as DNMT1, DNMT3A-3L, DNMT3B-3L and BCDIN3D (an RNA-methyltransferase) were assessed.
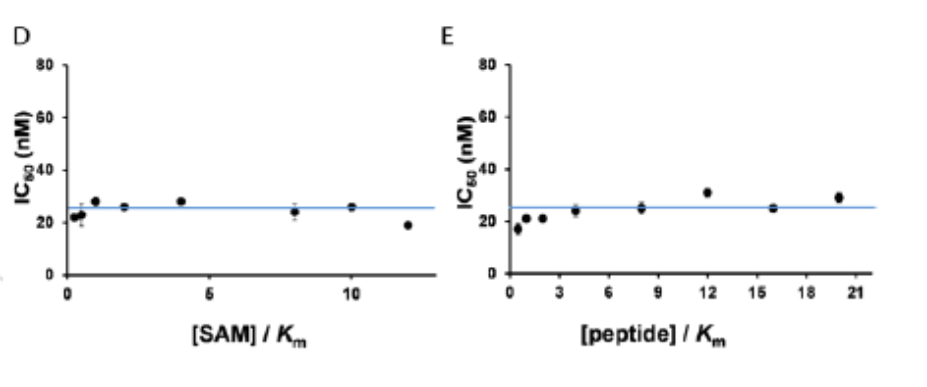
(D) IC50 values were determined at saturating concentration of peptide substrate (1.5 µM) and SAM concentrations equal to 0.25, 0.5, 1, 2, 4, 8, 10 and 12x of the Km for SAM. No change in IC50 values at varying SAM concentrations was consistent with a noncompetitive pattern of inhibition with respect to SAM. (E) To determine the competition with peptide, the SAM concentration was kept at saturation (70 µM) and IC50 values were determined at different peptide concentrations (0.5, 1, 2, 4, 8, 12, 16, and 20 ×Km). A similar noncompetitive pattern was observed for SGC707 with respect to peptide confirming the allosteric mode of inhibition.
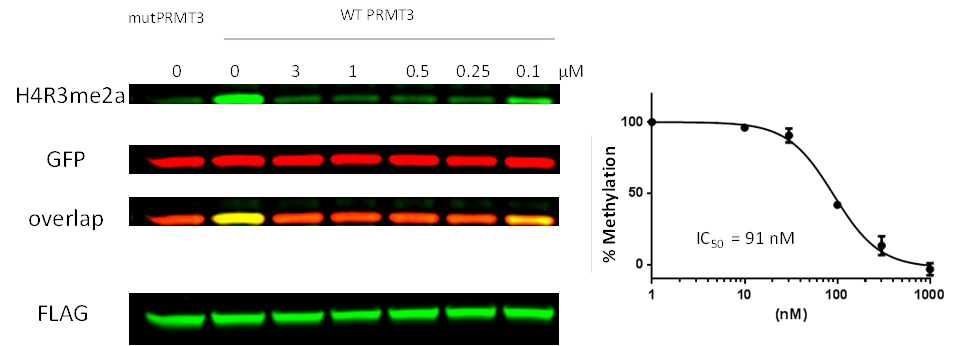
Western blot analysis of H4R3me2a levels. HEK293 cells were co-transfected with FLAG tagged PRMT3 (WT) or its catalytically dead mutant E335Q (Mut) and treated with different concentrations of SGC707, as indicated. Total cell lysates were collected 24 h post inhibitor treatment and analysed for H4R3me2a levels. The total levels of exogenous histone H4 and overexpressed PRMT3 were determined with anti-GFP, anti-H4 and anti-FLAG antibodies, respectively
Kaniskan, H. Ü., Szewczyk, M. M., Yu, Z., Eram, M. S., Yang, X., Schmidt, K., Luo, X., Dai, M., He, F., Zang, I., Lin, Y., Kennedy, S., Li, F., Dobrovetsky, E., Dong, A., Smil, D., Min, S.-J., Landon, M., Lin-Jones, J., Huang, X.-P., Roth, B. L., Schapira, M., Atadja, P., Barsyte-Lovejoy, D., Arrowsmith, C. H., Brown, P. J., Zhao, K., Jin, J. and Vedadi, M. (2015), A Potent, Selective and Cell-Active Allosteric Inhibitor of Protein Arginine Methyltransferase 3 (PRMT3).
Angew. Chem.. doi: 10.1002/ange.201412154